Escherichia coli (E. coli)
The encyclopedic reference for clinical microbiology, genomics, pathotypes, environmental indicators, and antimicrobial resistance. Discovered by Theodor Escherich in 1885.

Complete Taxonomy & Genomic Architecture
| Domain | Bacteria |
|---|---|
| Phylum | Pseudomonadota (formerly Proteobacteria) |
| Class | Gammaproteobacteria |
| Order | Enterobacterales |
| Family | Enterobacteriaceae |
| Genus | Escherichia |
| Species | E. coli |
| Type Strain | ATCC 11775 / NCTC 9001 |
Genomics & Evolution
E. coli possesses a highly dynamic genome, making it incredibly adaptable. Plasmids and bacteriophages drive rampant horizontal gene transfer (HGT).
- Genome Size: Approx. 4.6 to 5.5 Megabase pairs (Mbp), encoding ~4,000 to 5,500 genes depending on the strain.
- Core Genome vs. Pan-genome: Only about 20% of the genome is conserved across all strains (core). The pan-genome is massive, exceeding 90,000 unique gene families.
- Plasmids: Can carry the F-plasmid (fertility/conjugation), R-plasmids (antibiotic resistance), and Col-plasmids (bacteriocins).
- Generation Time: Under optimal conditions (37°C in rich media), it replicates every 20 minutes.
Morphology & Antigenic Structure
E. coli is a Gram-negative, facultative anaerobic, non-sporulating straight bacillus (rod) measuring approximately 1.0–3.0 µm in length and 0.4–0.7 µm in diameter. It is universally categorized using the Kauffmann-White classification system based on surface antigens.
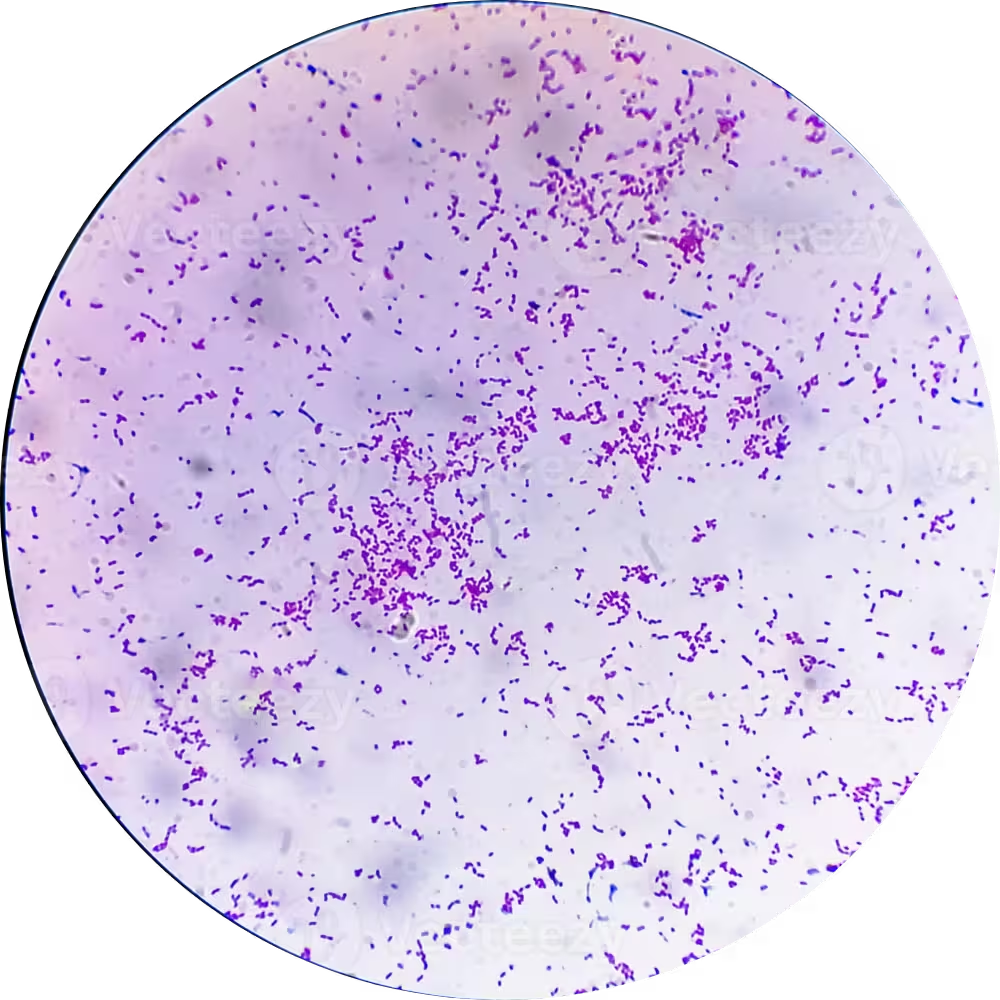
O Antigen (Somatic)
Comprises the outer polysaccharide portion of the Lipopolysaccharide (LPS) in the outer membrane. It is heat-stable and functions as a massive endotoxin (Lipid A). There are over 180 distinct O serogroups (e.g., O157, O104).
H Antigen (Flagellar)
Made of the protein flagellin. It is heat-labile. E. coli uses peritrichous flagella for swimming and swarming motility. There are ~53 identified H antigens (e.g., H7).
K & F Antigens
K Antigen (Capsule): An acidic polysaccharide microcapsule that inhibits phagocytosis and complement lysis (e.g., K1 antigen in neonatal meningitis).
F Antigen (Fimbriae/Pili): Proteinaceous appendages critical for adhesion to host epithelia (e.g., Type 1 pili, P-fimbriae).
Comprehensive Culture Media Profiles
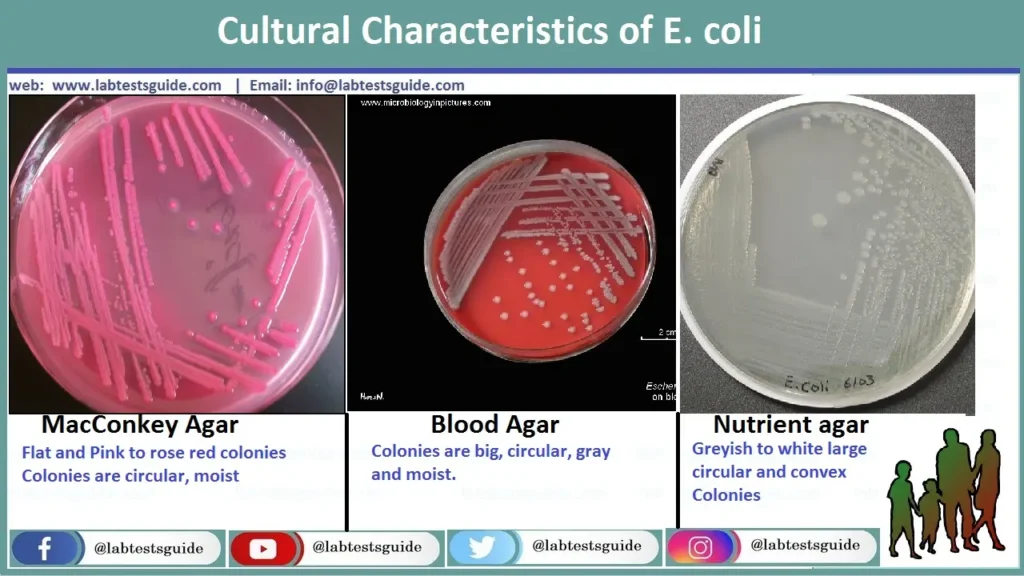
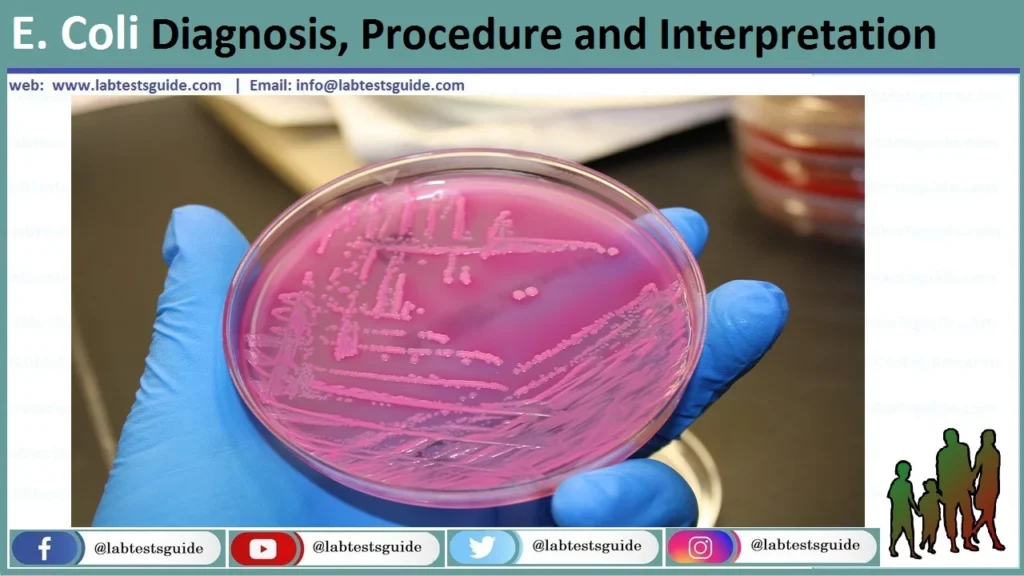
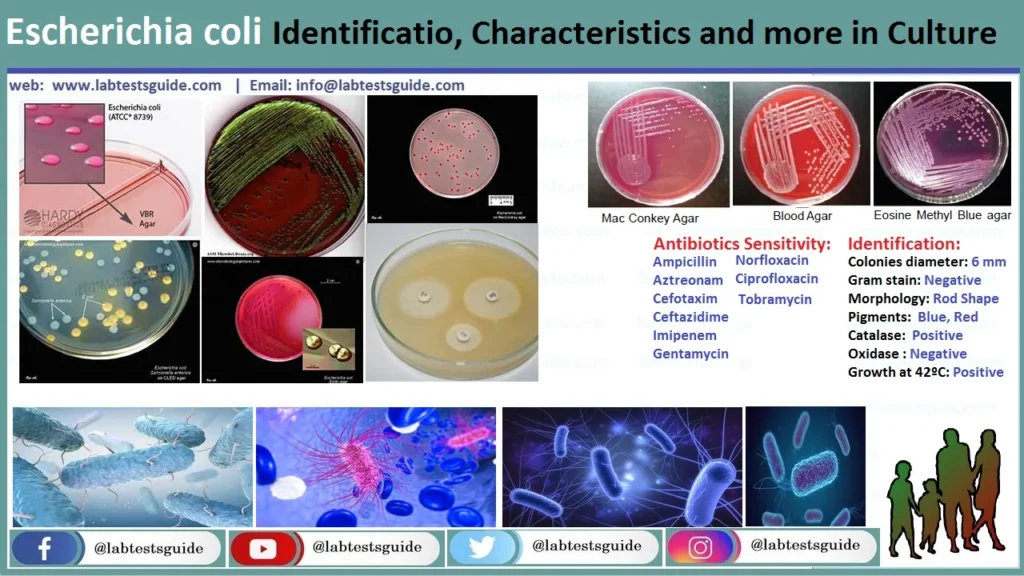
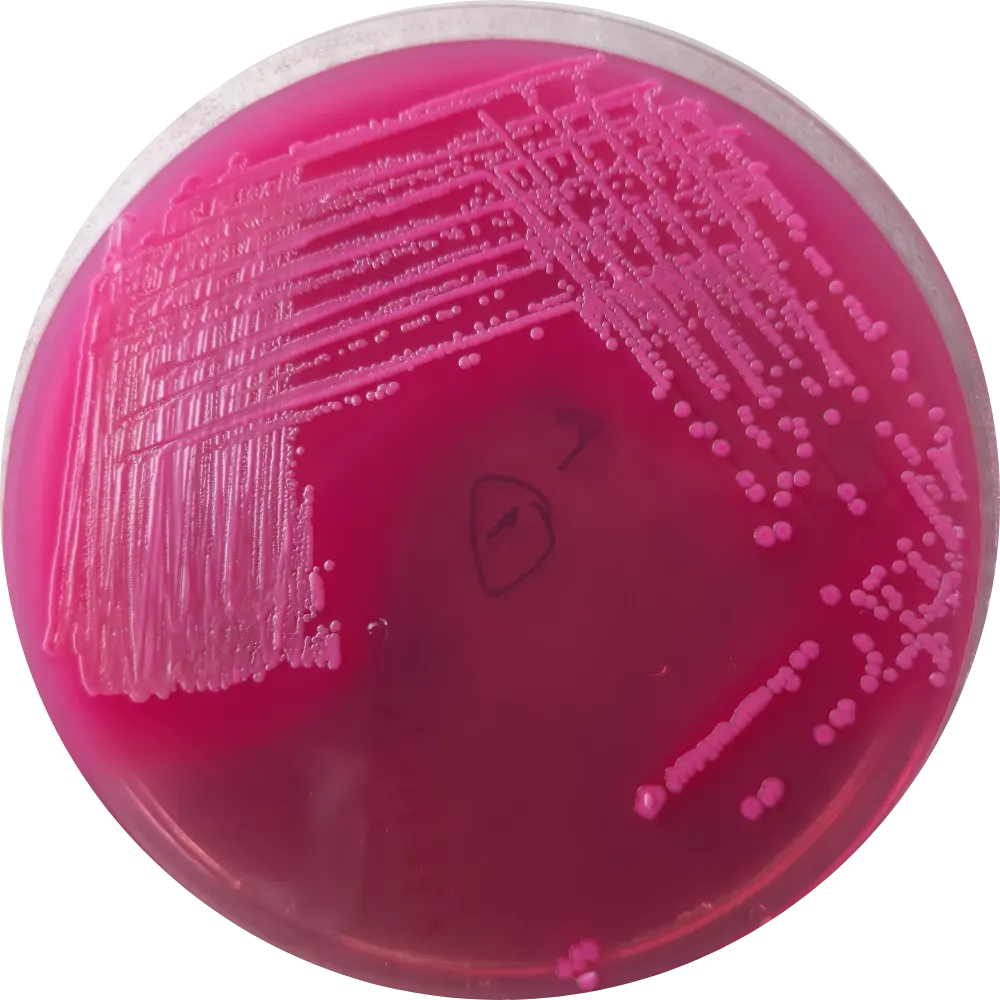
MacConkey (MAC)
Result: Bright pink/red colonies with a hazy bile precipitate zone.
Mech: Rapid lactose fermentation drops pH, turning neutral red indicator pink.
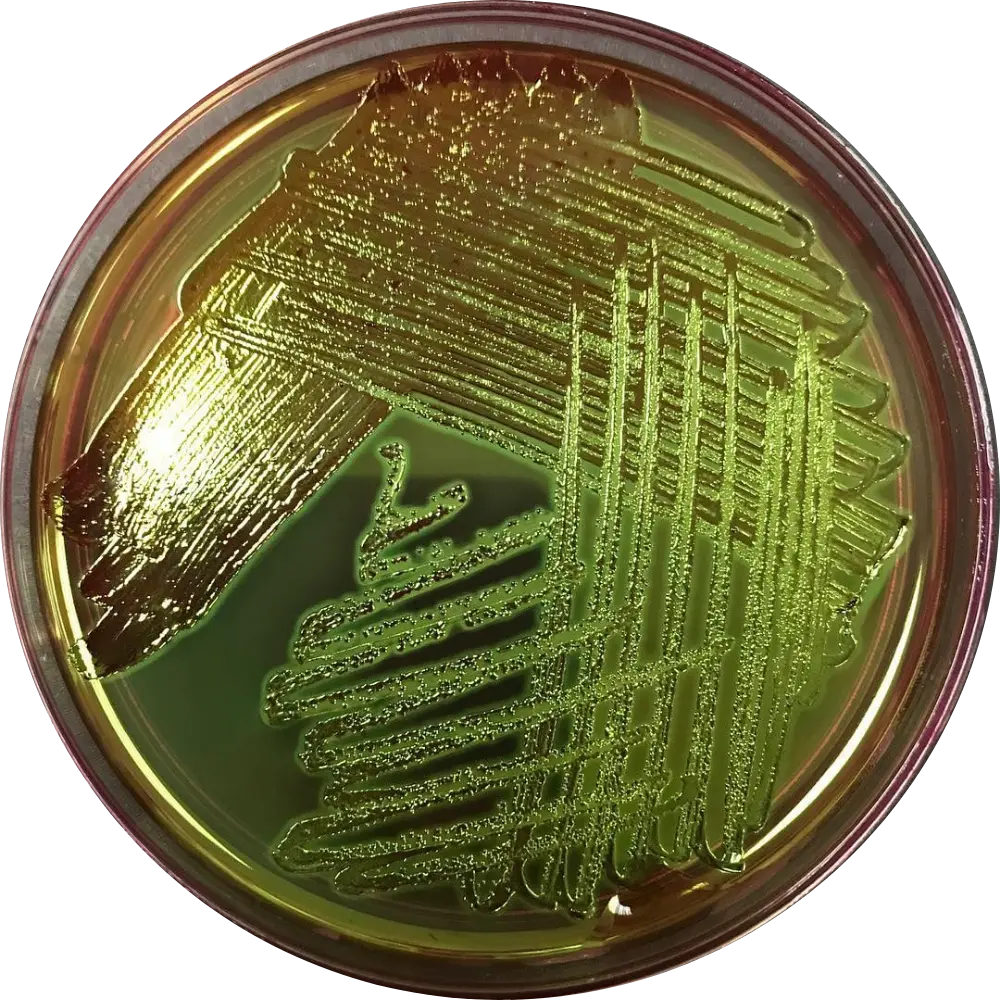
EMB Agar
Result: Dark-centered colonies with a striking Green Metallic Sheen.
Mech: Mixed-acid pathway produces massive acid, precipitating Eosin and Methylene Blue dyes.
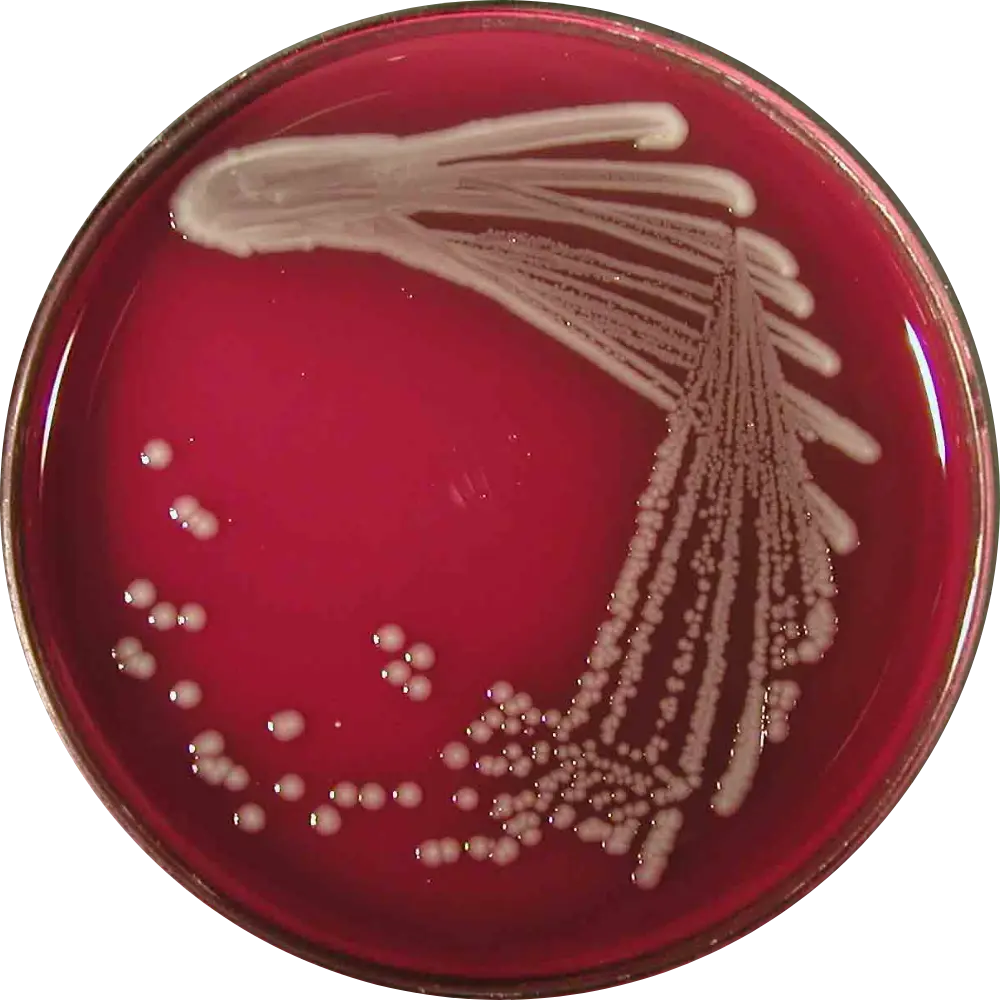
Blood Agar (BAP)
Result: Large, greyish colonies.
Note: Commensals are Gamma-hemolytic. Uropathogenic/Extraintestinal strains are often Beta-hemolytic (producing alpha-hemolysin).
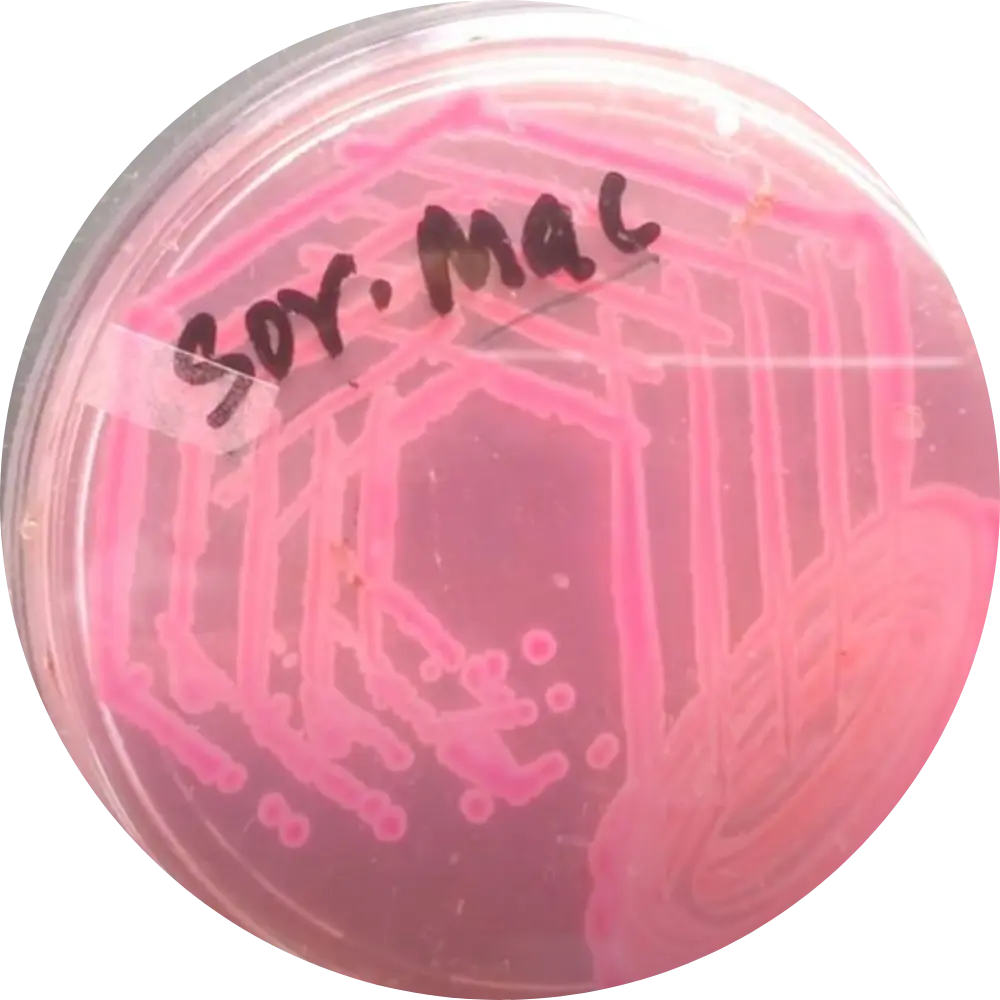
Sorbitol MAC (SMAC)
Result: Colorless colonies for EHEC O157:H7.
Mech: Unlike 95% of E. coli, O157:H7 is sorbitol-negative. Regular strains appear pink.
CLED Agar
Result: Large yellow colonies.
Use: Standard for urine cultures. Lacks electrolytes to inhibit Proteus swarming.
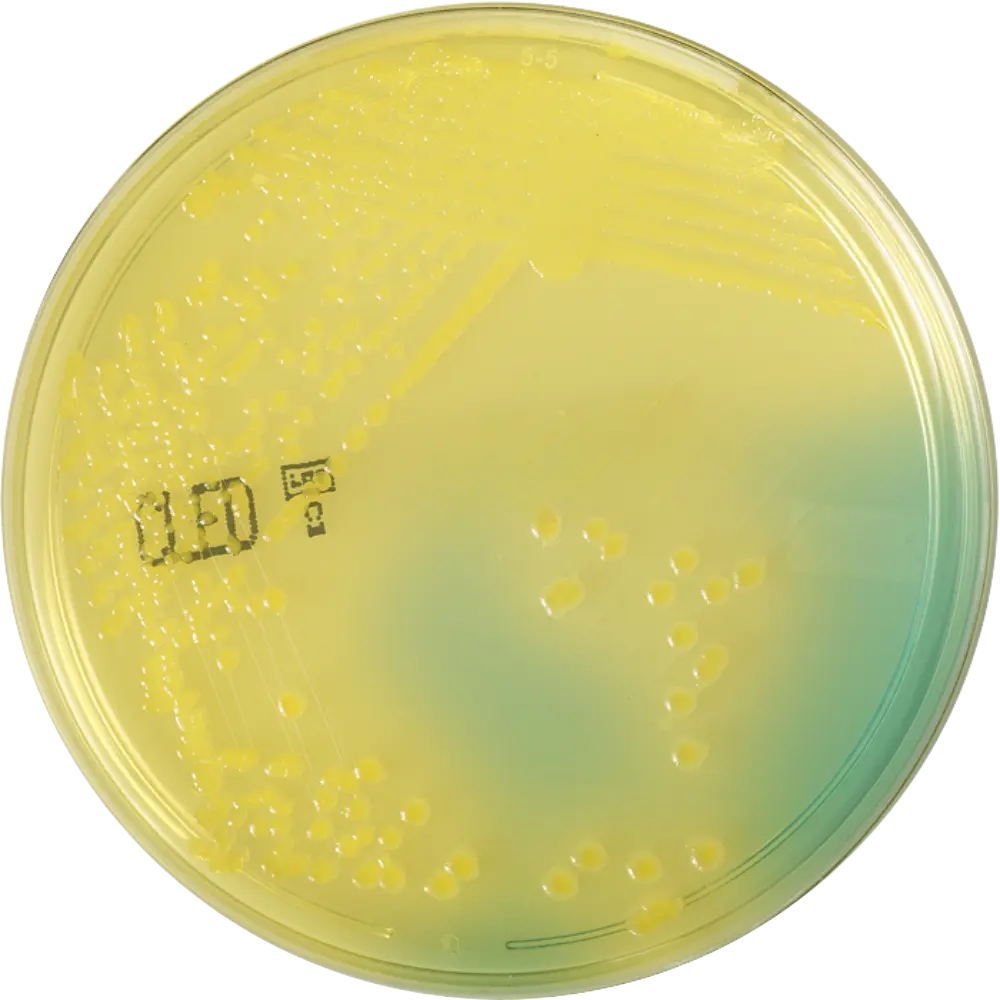
CHROMagar™ E. coli
Result: Dark blue/purple colonies.
Mech: Detects specific enzymes (like beta-glucuronidase) using proprietary chromogenic substrates.
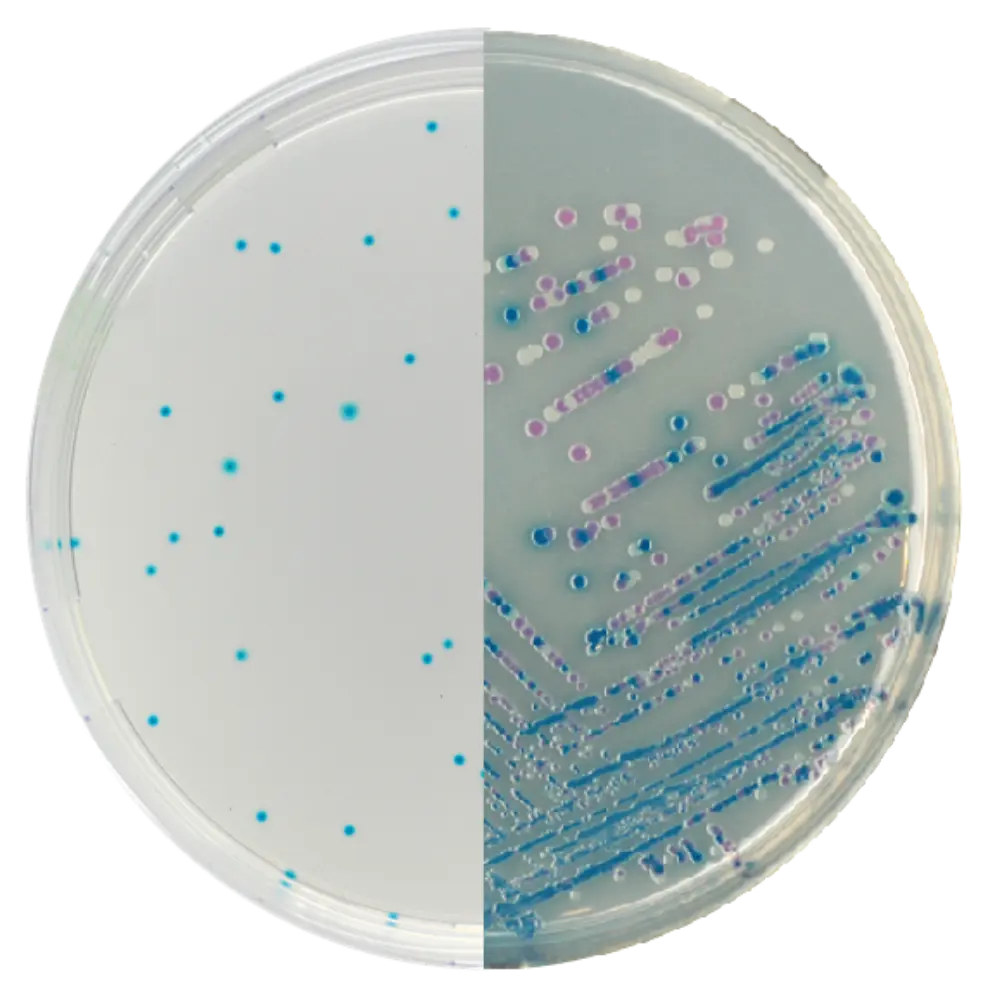
XLD Agar
Result: Large yellow colonies.
Mech: Ferments lactose/xylose. Differentiates from Salmonella (which has black centers due to H2S).
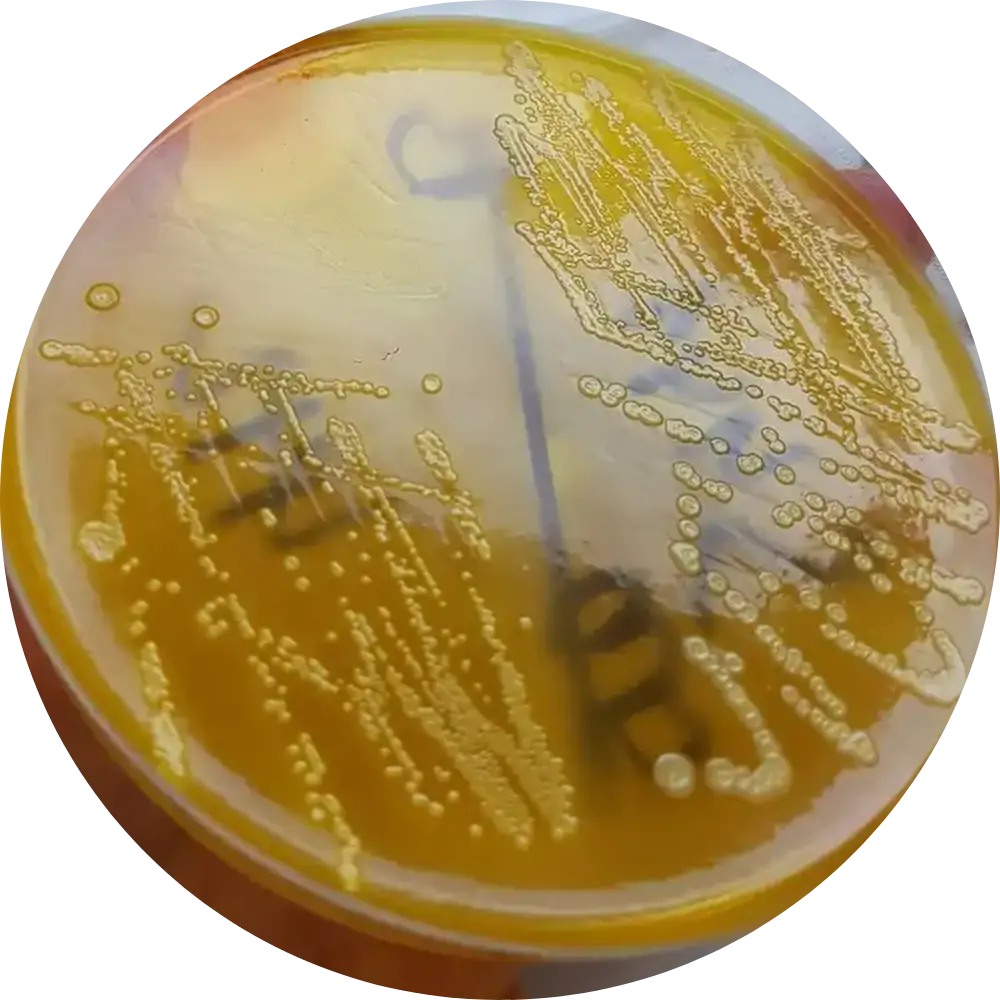
SS / HE Agar
Result: Escherichia coli forms pink/red lactose-fermenting colonies on SS agar and yellow/orange colonies on HE agar, with no H₂S production (no black centers).
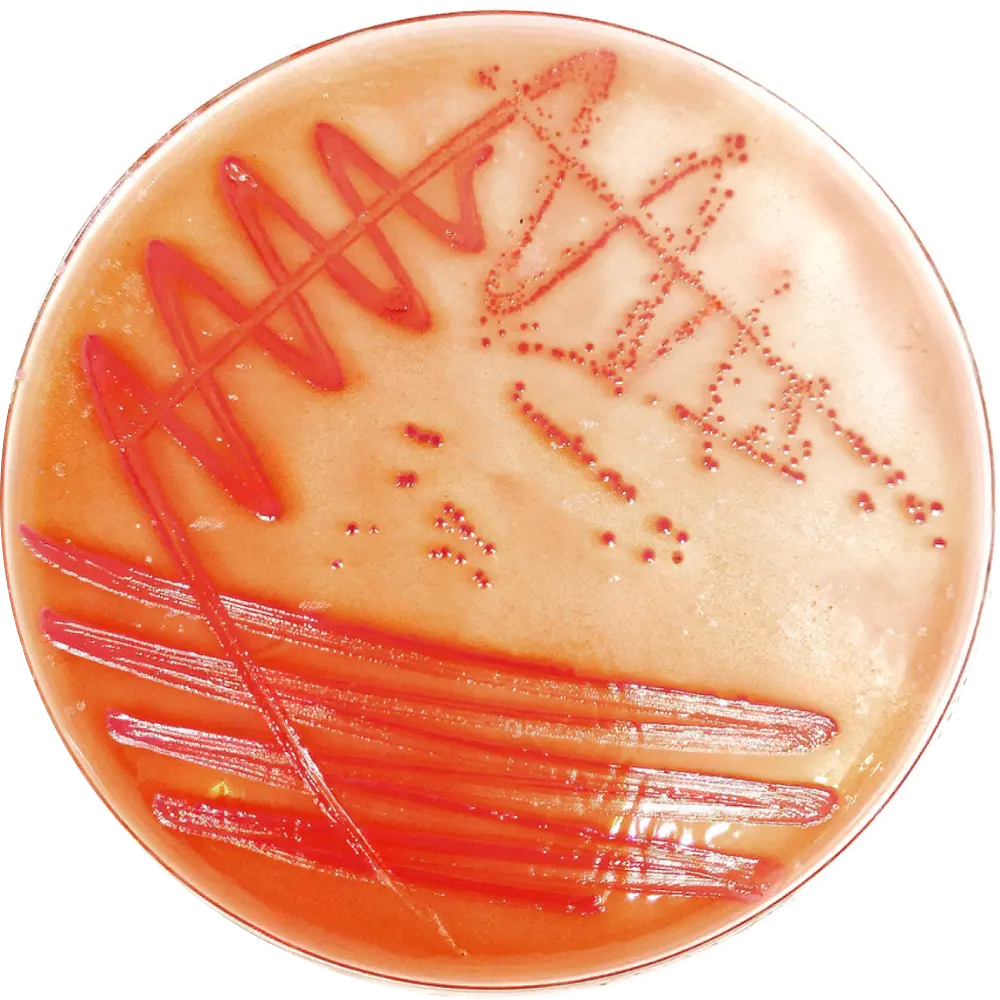
Complete Biochemical Diagnostics
| Test | Result | Biochemical Mechanism & Observation |
|---|---|---|
| Indole / Spot Indole | Positive | Tryptophanase enzyme cleaves amino acid tryptophan. Turns red with Kovac’s reagent (or blue with DMACA on spot test). |
| Methyl Red (MR) | Positive | Utilizes the mixed-acid fermentation pathway, creating stable acids that drop pH below 4.4 (turns red). |
| Voges-Proskauer (VP) | Negative | Does not use the butylene glycol pathway; no acetoin is produced. |
| Citrate | Negative | Lacks citrate permease; cannot transport citrate into the cell. Agar remains green. |
| Triple Sugar Iron (TSI) | A/A, Gas+, H2S- | Ferments glucose, lactose, and sucrose (Yellow slant/Yellow butt). Produces CO2/H2 gas (cracks agar). Lacks thiosulfate reductase (no black H2S precipitate). |
| Urease | Negative | Cannot hydrolyze urea into ammonia. Media stays yellow/orange (differentiates from Proteus/Klebsiella). |
| Decarboxylases (Moeller) | LDC+, ODC(V), ADH- | Positive for Lysine Decarboxylase (LDC). Variable for Ornithine (ODC). Negative for Arginine Dihydrolase (ADH). |
| MUG Test | Positive | Produces β-glucuronidase, which cleaves MUG to release a compound that fluoresces brilliant blue under UV light. (Note: O157:H7 is MUG negative). |
| Nitrate Reduction | Positive | Reduces Nitrates (NO3) to Nitrites (NO2), a hallmark of all Enterobacteriaceae. |
| Cytochrome Oxidase | Negative | Lacks cytochrome C oxidase (differentiates from Pseudomonas). |
Mechanisms of Pathogenicity (Virulence Factors)
Commensal E. coli transitions to a pathogen through the acquisition of pathogenicity islands (PAIs), plasmids, and bacteriophages encoding destructive proteins.
A. Adhesins (Pili & Fimbriae)
- Type 1 Fimbriae: Bind to mannose residues. Present in almost all strains. Associated with lower UTIs (cystitis).
- P-Fimbriae (Pap pili): Bind to the P blood group antigen (globobiose) on uroepithelial cells. Essential for upper UTIs (pyelonephritis) as they resist urine flushing.
- Bundle-Forming Pili (BFP): Used by EPEC to cause microcolony formation on the intestinal epithelium.
B. Exotoxins & Endotoxins
- Shiga Toxins (Stx1, Stx2): AB5 toxins encoded by lambdoid phages in EHEC. They cleave the 28S rRNA of the 60S ribosomal subunit, halting host protein synthesis and causing cell death (leading to HUS).
- Heat-Labile (LT) & Heat-Stable (ST) Enterotoxins: Produced by ETEC. LT increases cAMP, ST increases cGMP. Both cause massive efflux of water and electrolytes into the gut lumen (watery diarrhea).
- Alpha-Hemolysin (HlyA): Forms pores in host cell membranes, liberating nutrients and iron. Common in UPEC.
- Lipid A (Endotoxin): Part of the LPS. Triggers massive release of TNF-alpha and IL-1 when bacteria lyse, causing Septic Shock and SIRS.
C. Iron Acquisition (Siderophores) & Secretion Systems
- Enterobactin & Aerobactin: High-affinity iron-chelating compounds secreted by E. coli to steal iron from host transferrin and lactoferrin, essential for survival in blood and urine.
- Type III Secretion System (T3SS): A molecular “syringe” used by EPEC and EHEC to inject effector proteins (like Tir) directly into host intestinal cells, manipulating host actin to form Attaching & Effacing (A/E) pedestal lesions.
Clinical Pathotypes & Associated Diseases
Intestinal (Diarrheagenic) Pathotypes
ETEC Traveler’s Diarrhea
Epi: Leading cause of traveler’s and weanling diarrhea in developing nations.
Pathogenesis: Non-invasive. Attaches via CFAs. Secretes LT/ST toxins causing hypersecretion of chloride.
Symptoms: Profuse, watery, cholera-like diarrhea. No inflammation/fever.
EPEC Infantile Diarrhea
Epi: Endemic infant diarrhea.
Pathogenesis: Binds via BFP. Uses T3SS to inject Tir, causing actin rearrangement and microvilli destruction (Attaching/Effacing lesions). Malabsorption follows.
Symptoms: Prolonged watery diarrhea with mucus.
EHEC / STEC Bloody Diarrhea / HUS
Epi: O157:H7 (undercooked beef, raw milk, unpasteurized cider).
Pathogenesis: Produces Shiga-toxins (Stx) causing capillary microthrombi and endothelial damage in gut and kidneys.
Symptoms: Hemorrhagic colitis (severe bloody diarrhea, NO fever). Complicated by Hemolytic Uremic Syndrome (HUS) in 10% of cases.
EAEC Persistent Diarrhea
Pathogenesis: “Stacked-brick” autoaggregation via AAF fimbriae. Forms thick mucus biofilm and secretes EAST1 cytotoxin.
Symptoms: Chronic diarrhea (>14 days) in pediatric and HIV/AIDS populations.
EIEC Dysentery
Pathogenesis: Phenotypically identical to Shigella. Relies on the pInv plasmid to invade enterocytes, escape the phagosome, and spread laterally via actin tails.
Symptoms: Fever, cramps, tenesmus, and bloody/mucoid diarrhea loaded with neutrophils.
DAEC Pediatric Diarrhea
Pathogenesis: Diffuse adherence covering the entire cell surface. Stimulates signal transduction causing elongated microvilli that wrap the bacteria.
Symptoms: Watery diarrhea in children aged 1-5.
Extra-Intestinal Pathotypes (ExPEC)
UPEC (Uropathogenic)
Responsible for 80-90% of all community-acquired UTIs. Uses Type 1 pili to colonize the bladder (cystitis) and P-fimbriae to ascend to the kidneys (pyelonephritis). Often produces hemolysins.
NMEC (Neonatal Meningitis)
Causes ~20% of neonatal meningitis cases. Highly dependent on the K1 capsular antigen, a sialic acid polymer that mimics human neural cell adhesion molecules, evading host defenses.
APEC (Avian Pathogenic)
Causes severe respiratory and systemic diseases in poultry (Colibacillosis). Zoonotic potential: shares identical virulence plasmids with human UPEC and NMEC strains, causing foodborne extra-intestinal infections.
Comprehensive Laboratory Diagnostics
Step 1: Pre-Analytical Collection
- Urine: Mid-stream clean-catch (MSCC), straight catheter, or suprapubic aspirate. Use boric acid transport tubes if plating is delayed >2 hours. Quantitative threshold: >10⁵ CFU/mL (or >10³ for catheterized).
- Stool: Fresh feces or rectal swabs in Cary-Blair transport media (preserves delicate EHEC/Campylobacter).
- Blood/CSF: Inoculate aerobic and anaerobic BACTEC/BacT/ALERT bottles immediately.
Step 2: Analytical Identification (Traditional & Molecular)
- Culture & Biochemistry: Plate to BAP, MAC, and EMB. Confirm LF colonies with spot indole (positive). If O157:H7 is suspected, plate to SMAC and look for colorless colonies.
- MALDI-TOF MS: Matrix-Assisted Laser Desorption/Ionization Time-of-Flight Mass Spectrometry. The modern gold standard. Identifies E. coli in 2 minutes by analyzing ribosomal protein signatures against a massive database.
- Multiplex PCR / FilmArray: Syndromic GI or Blood culture panels can detect E. coli DNA and specify pathotype virulence genes (e.g., detecting stx1, stx2, eae, pInv) directly from raw stool or blood in 1 hour.
Step 3: Serotyping (Kauffmann-White)
Used primarily by public health labs (CDC/State labs) during outbreaks. Involves slide agglutination tests utilizing specific antisera to identify exact O and H antigens (e.g., confirming a non-sorbitol fermenter is indeed O157 and H7 positive).
Antimicrobial Resistance (AMR) & Treatment
Standard Pharmacotherapy
AST must be performed via Broth Microdilution (MIC) or Kirby-Bauer disc diffusion.
- Uncomplicated Cystitis (UTI): Nitrofurantoin, Fosfomycin, or Trimethoprim-sulfamethoxazole (TMP-SMX).
- Pyelonephritis / Septicemia: Intravenous Fluoroquinolones (Ciprofloxacin) or 3rd-Generation Cephalosporins (Ceftriaxone).
- Gastroenteritis: Primarily supportive (Oral Rehydration Therapy). Antibiotics only for severe traveler’s diarrhea (Azithromycin).
Global AMR Threats
E. coli rapidly exchanges R-plasmids via conjugation, leading to critical resistance paradigms:
- ESBL Producers: Extended-Spectrum Beta-Lactamases (CTX-M, TEM, SHV enzymes) hydrolyze penicillins and all cephalosporins. Treatment requires Carbapenems (Meropenem).
- CRE (Carbapenem-Resistant Enterobacteriaceae): Produce enzymes like KPC or NDM-1. Resistant to almost all beta-lactams.
- MCR-1 Gene: Plasmid-mediated resistance to Colistin (Polymyxin E), a highly toxic drug of last resort. MCR-1 alters the lipid A target, creating pan-resistant “nightmare bacteria.”
Environmental Health, Water Quality & Biotechnology
Universal Indicator Organism
Because E. coli cannot survive long outside the host, its presence in environmental water strictly indicates recent fecal contamination. EPA regulations mandate 0 CFU/100mL in drinking water.
- Colilert System: Uses ONPG (detects total coliforms – turns yellow) and MUG (detects E. coli specifically – fluoresces blue under UV).
- Membrane Filtration: 100mL of water is pulled through a 0.45µm filter and placed on m-Endo or m-TEC agar.
- Most Probable Number (MPN): Statistical estimation using multiple fermentation tubes.
The Ultimate Model Organism
Strains like K-12 and BL21 are harmless biosafety level 1 organisms that have driven modern biology.
- Historical Milestones: Used to discover bacterial conjugation (Joshua Lederberg) and gene regulation via the lac operon (Jacob & Monod).
- Recombinant DNA Bio-factory: Using heat-shock transformation or electroporation, human genes are inserted into E. coli plasmids. It currently produces massive global supplies of Human Insulin, Human Growth Hormone (HGH), and industrial enzymes.
Rapid Review: MLS / ASCP Board Pearls
Biochemicals
IMViC: + + – –
TSI: A/A, Gas+, H2S-
Urease: Negative
Oxidase: Negative
Agar Appearances
MAC: Bright Pink (LF)
EMB: Green Metallic Sheen
SMAC: Colorless (O157)
CLED: Yellow
Key Pathotypes
EHEC: Shiga Toxin, HUS
UPEC: P-fimbriae (UTI)
NMEC: K1 Capsule (Meningitis)
ETEC: LT/ST Toxin
Critical Rules
Never give antibiotics for O157:H7.
MUG test differentiates generic E. coli (+) from O157 (-).