Urine Culture Test Defination Purpose, Types, Procedure, Result Interpretation and moreUpdated 2026
Comprehensive Reference for Lab Professionals & Students

💡 Definition & Purpose
Definition: A urine culture is a laboratory test to detect and identify bacteria or other microorganisms (like fungi) in urine by inoculating a sample onto a culture medium. It is the gold standard for diagnosing urinary tract infections (UTIs) .
Purpose: To diagnose UTIs , identify the specific causative organism, and perform antibiotic susceptibility testing (AST) to guide effective treatment. It’s also used to monitor treatment efficacy and screen for asymptomatic bacteriuria in high-risk patients (e.g., pregnant women).
🩺 When is it Ordered?
Clinical Indications
- Symptoms of UTI (dysuria, frequency, urgency).
- Suspected pyelonephritis (fever, flank pain).
- Screening during pregnancy.
- Patients with indwelling catheters.
- Recurrent or complicated UTIs.
- Pre-surgical urological evaluation.
Common Symptoms
- Pain or burning during urination (dysuria).
- Frequent urination (frequency).
- Sudden urge to urinate (urgency).
- Cloudy, foul-smelling, or bloody urine .
- Lower abdominal pain or pressure.
- Fever, chills, nausea (if kidney involved).
🧪 Specimen Collection & Handling
Collection Methods
- Clean-Catch Midstream: Standard method. Patient cleans genital area, starts urinating, then collects mid-portion. How to collect ?
- Catheterized Specimen: For patients unable to void or to avoid contamination.
- Suprapubic Aspiration: Gold standard for infants or when anaerobes are suspected. Bypasses urethra.
- Urine Collection Bag: Only for infants; high contamination rate.
Sample Stability & Transport
- Room Temp: ≤ 2 hours (delay allows bacterial multiplication).
- Refrigerated (2-8°C): Up to 24 hours. Use boric acid preservative tubes for longer transport.
- Frozen: Not recommended for culture.
- Container: Sterile, leak-proof, wide-mouth container.
Rejection Criteria
- Unlabeled or mislabeled specimen.
- Leaking container.
- Transport delay >2 hours (unrefrigerated).
- Collection from urine bag in adults.
- Insufficient quantity (<3 mL).
🔬 Laboratory Procedure

Inoculation & Incubation
- Mix urine sample gently.
- Use a calibrated loop (1 µL or 10 µL) to inoculate:
- Blood Agar (BA) : For fastidious organisms and hemolysis observation.
- MacConkey Agar (MAC) : Selective for Gram-negatives, differentiates lactose fermenters.
- CLED Agar : Supports both Gram-positives and negatives, prevents Proteus swarming.
Incubation: Aerobically at 35-37°C for 18-24 hours. If no growth, re-incubate for another 24 hours.
Colony Count & Interpretation
Number of colonies = CFU/mL. A 1 µL loop (0.001 mL) means 1 colony = 1,000 CFU/mL.
| CFU/mL | Interpretation (Symptomatic) |
|---|---|
| ≥ 100,000 | Significant bacteriuria (infection). |
| 10,000 – 100,000 | Suspicious; correlate with symptoms. |
| < 10,000 | Likely contamination; repeat if symptomatic. |
| Mixed flora (>2 types) | Contamination likely. |
🦠 Common & Uncommon Uropathogens
Common Isolates
- Escherichia coli (80-85% of community-acquired UTIs)
- Klebsiella pneumoniae
- Staphylococcus Saprophyticus (young women)
- Enterococcus faecalis
- Proteus mirabilis (associated with kidney stones)
- Pseudomonas Aeruginosa (healthcare-associated)
- Enterobacter Cloacae , Citrobacter spp.
- Group B Streptococcus (pregnancy)
Uncommon/Special Pathogens
- Mycoplasma hominis, Ureaplasma urealyticum (require special media)
- Neisseria gonorrhoeae (sexually transmitted)
- Chlamydia trachomatis (not detectable by routine culture)
- Candida spp. (yeast, especially in immunocompromised)
- Mycobacterium tuberculosis (requires special AFB culture)
- Acinetobacter, Burkholderia, Corynebacterium urealyticum
📋 Pathogen Identification Profiles
Key cultural and biochemical characteristics for definitive identification.

| Organism | Gram Stain | Key Media Growth | Key Biochemicals | Notes |
|---|---|---|---|---|
| E. coli | G- rod | Pink colonies on MAC (Lac+) | Indole (+), Motile (+), Oxidase (-) | Most common UTI pathogen |
| Klebsiella pneumoniae | G- rod | Pink, mucoid on MAC (Lac+) | Citrate (+), Urease (+), Non-motile | Prominent polysaccharide capsule |
| Proteus mirabilis | G- rod | Swarming on BA, clear on MAC (Lac-) | Urease (+), H2S (+), PAD (+), Motile | Associated with struvite stones |
| Pseudomonas aeruginosa | G- rod | Non-lactose fermenter, grape-like odor, blue-green pigment | Oxidase (+), Motile, Pyocyanin (+) | Intrinsically resistant to many drugs |
| Enterococcus faecalis | G+ cocci (pairs/chains) | Small, gamma-hemolytic on BA, grows on Bile Esculin agar | Catalase (-), PYR (+), Esculin hydrolyzed | Intrinsic cephalosporin resistance |
| S. saprophyticus | G+ cocci (clusters) | Grows on MSA (yellow), white colonies | Catalase (+), Coagulase (-), Novobiocin Resistant | UTI in young women |
| S. aureus | G+ cocci (clusters) | Beta-hemolytic, yellow colonies on MSA | Catalase (+), Coagulase (+) | Less common, but can cause UTI |
| Candida albicans | Yeast (fungus) | Creamy colonies on BA or Sabouraud agar | Germ tube (+), Chlamydospore (+) | Risk factors: diabetes, antibiotics, catheters |
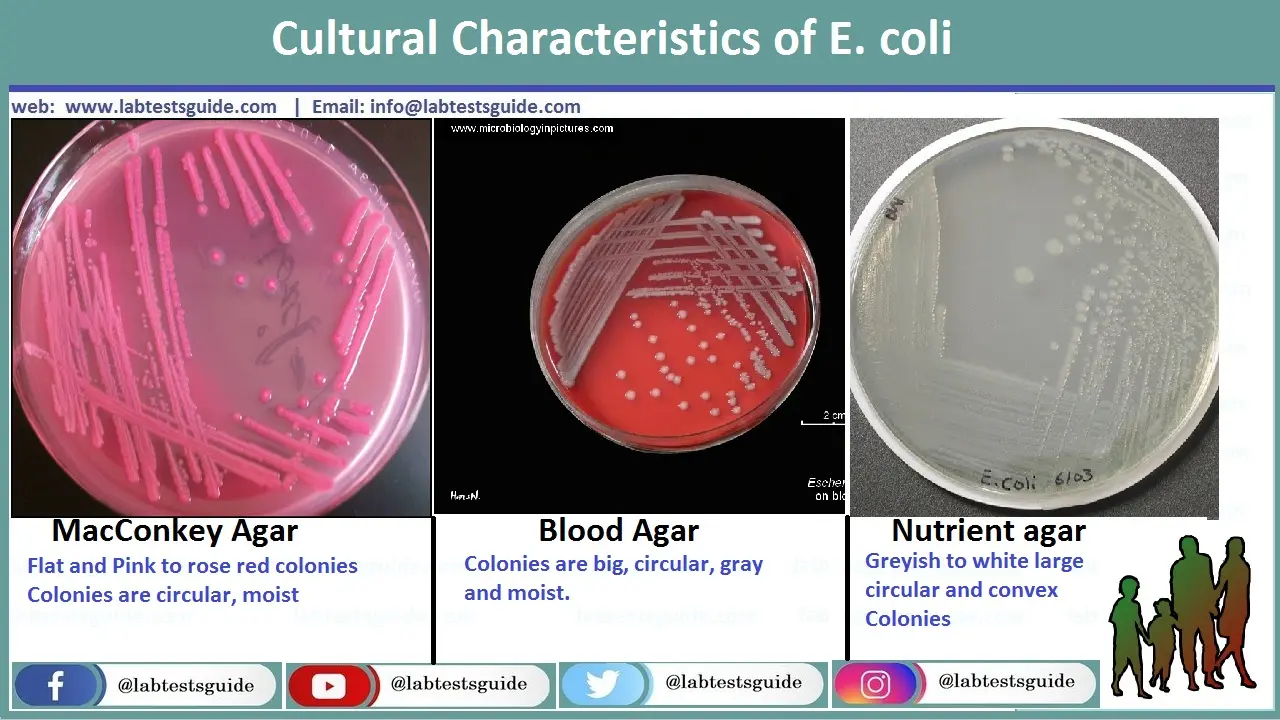
E. Coli on Blood, MacConkey and Neutrient Agar in Urine sample
Pink Colonies on MAC (Lac+), Gray and Moist Circular Big Colonies on Blood Agar, Grayish to White Circular and Convex Smooth and Rough Colonies on Neutrient Agar
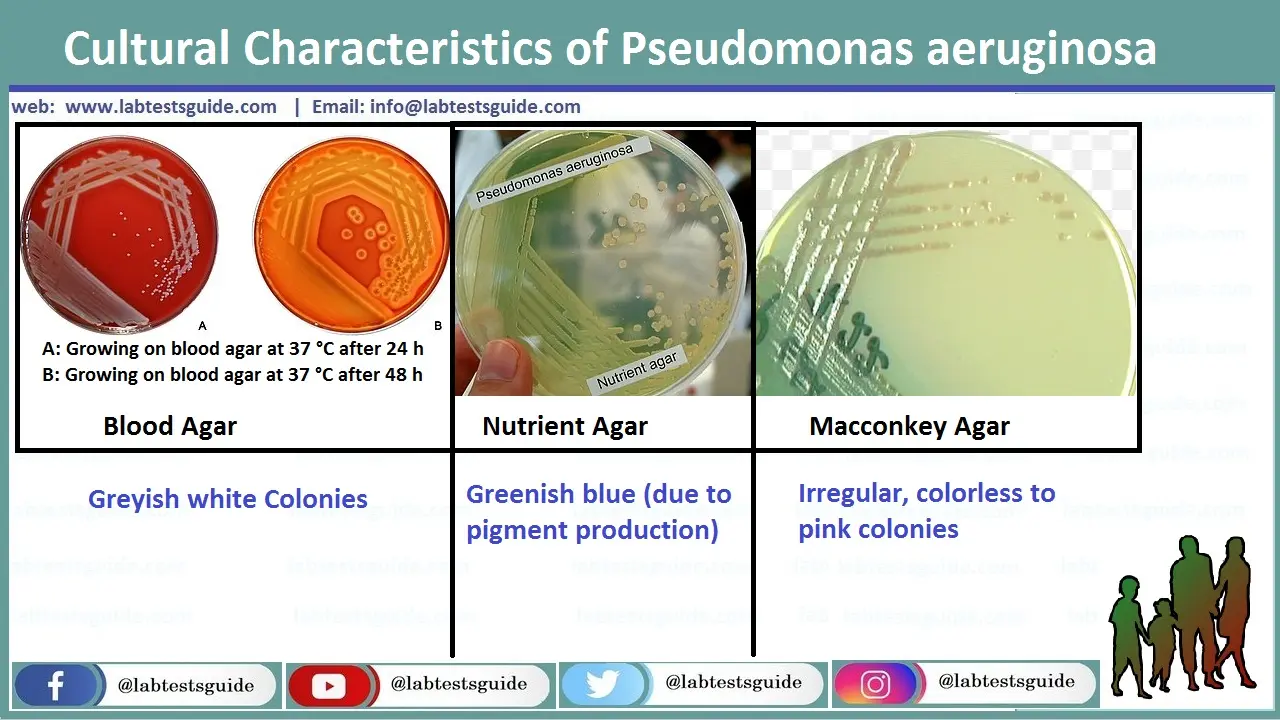

Pseudomonas Aeruginosa on Blood, MacConkey and Neutrient Agar in Urine Sample
Grayish white colonies on Blood Agar, Colorless to pink Irregular Colonies on MacConkey Agar, Greenish Blue Colonies on Neutrient Agar
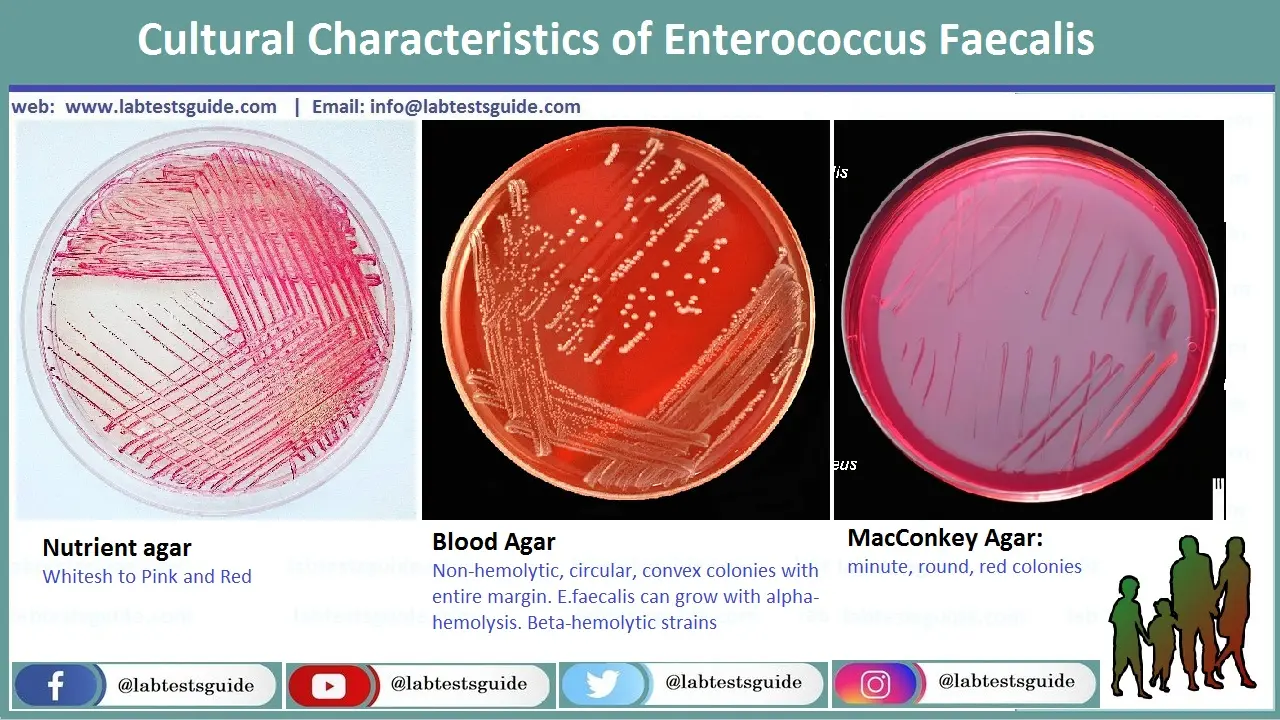
Enterococcus Faecalis on Blood, MacConkey and Neutrient Agar in Urine Sample
Minute, Rounded and red colonies on MacCOnkey Agar, Whitesh to Pink Colonien on Neutrient Agar, No-Hemolytic, Circular, Convex Colonies with entire margen on Blood Agar. E.Faecalis can grow with Alpha-Hemolysis, Beta-Hemolytic Strains
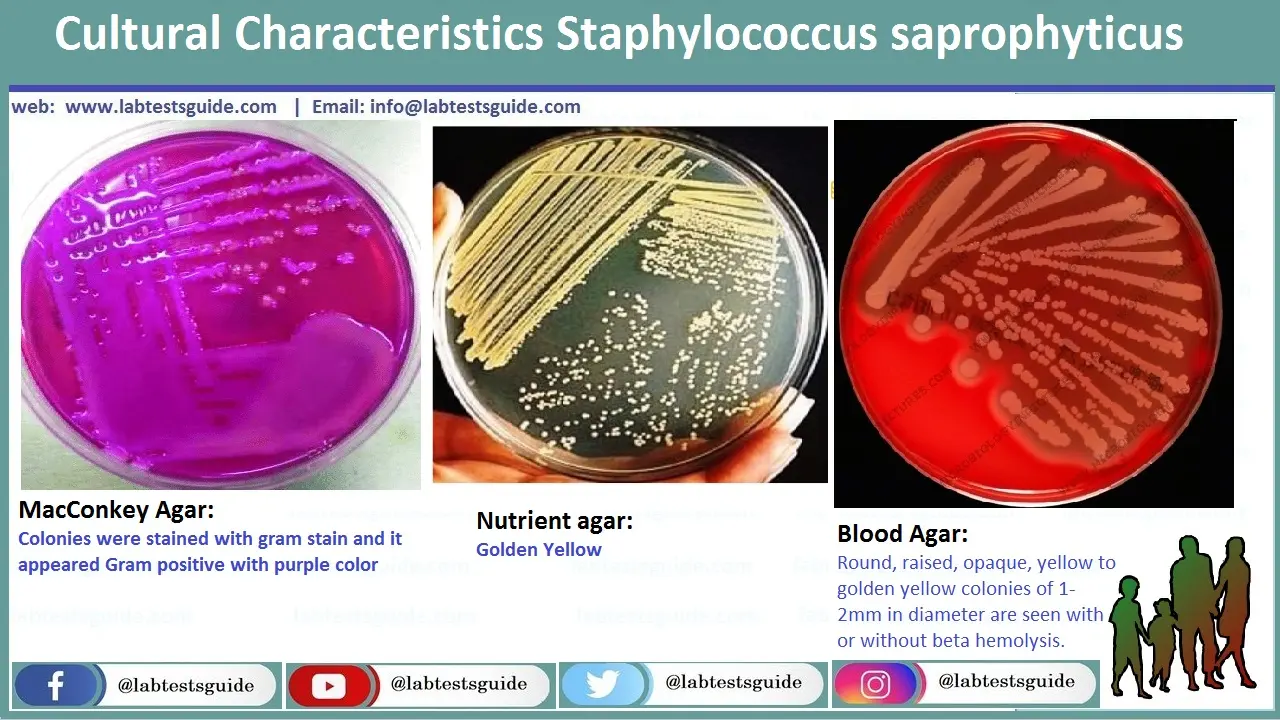
Staphylococcus Saprophyticus on Blood, MacConkey and Neutrient Agar in Urine Sample
Golden Yellow Colonies on Neutrient Agar, Colonies wuth Stained with gram Stain and it appeared Grap positive with Purple Color on MacConkey Agar, Rounded, Raised, apaque, Yellow to Golden Colonies of 1-2mm in diameter are seen with or without beta-hemolysis on Blood Agar
🧠 AI Diagnostic Matrix (Urine Edition)
Interactive tool to match lab findings with potential pathogens. Select observed traits to narrow down suspects.
🦠 Bacterial ID Matrix (Urine Edition)
AI-driven diagnostic matrix for Urinary Tract Pathogens.
💡 How to Use This Tool
- Step 1: Input Data – Select known lab parameters (Staining, Media, Biochemicals) from the panel.
- Step 2: Consult AI – Watch the AI panel for suggested tests to narrow down uropathogen suspects.
- Step 3: Finalize Report – Click “Verify & Finalize” on your best matching pathogen to generate a clinical summary.
- Step 5: Print & Reset – Print your verified report document, then click “Return to Matrix” for Proceed Again.
💊 Antibiotic Susceptibility Testing (AST)
Once a pathogen is identified, AST determines the most effective antibiotics. The Kirby-Bauer disk diffusion method is common. A lawn of bacteria is spread on Mueller-Hinton agar, and antibiotic-impregnated discs are placed. The zone of inhibition is measured and interpreted per CLSI/EUCAST standards.
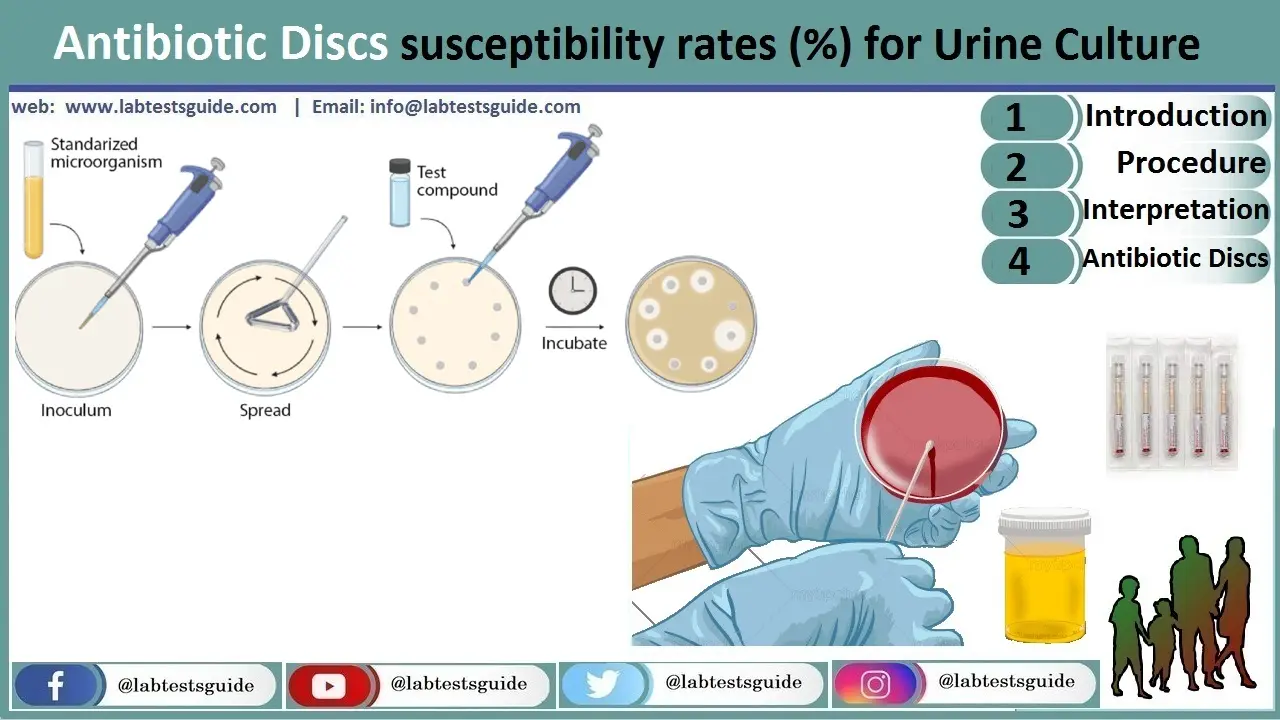

🧫 Common Antibiotic Discs for Urine Culture
Common Discs: Ampicillin, Amoxicillin-Clavulanate, Piperacillin-Tazobactam, Cefazolin, Ceftriaxone, Ceftazidime, Cefepime, Imipenem, Meropenem, Gentamicin, Amikacin, Tobramycin, Ciprofloxacin, Levofloxacin, Nitrofurantoin, Trimethoprim-Sulfamethoxazole, Fosfomycin, Vancomycin (for Gram-positives).

Susceptibility Patterns of Key Uropathogens (from studies)
| Antibiotic | E. coli (%S) | K. pneumoniae (%S) | P. aeruginosa (%S) | S. aureus (%S) | E. faecalis (%S) |
|---|---|---|---|---|---|
| Ampicillin | ~40-50% | 0% (intrinsic R) | 0% (R) | ~10-20% | >95% |
| Amoxicillin-Clav | ~70-80% | ~70-85% | 0% (R) | ~80-90% | >95% |
| Nitrofurantoin | >95% | ~50-70% | 0% (R) | >95% | >95% |
| Ciprofloxacin | ~70-85% | ~70-80% | ~70-80% | ~60-80% | ~50-70% |
| Trimethoprim-Sulfa | ~70-80% | ~70-80% | 0% (R) | >95% | Variable |
| Gentamicin | >90% | >90% | ~80-90% | >90% | Synergy only |
| Meropenem | >99% | >99% | >90% | N/A | N/A |
| Vancomycin | N/A | N/A | N/A | 100% | >95% |
Note: Resistance rates vary by region and over time. Always refer to local antibiograms. R = Resistant, S = Susceptible.
❓ Frequently Asked Questions (For Lab Pros & Students)
Urinalysis is a chemical/microscopic screen for abnormalities (like leukocytes, nitrites). Urine culture is a microbiological test to grow and identify specific bacteria and quantify them (CFU/mL).
To minimize contamination by normal flora from the urethra, skin, or genital area. The first portion of urine flushes out these contaminants.
Traditionally, ≥10^5 CFU/mL of a single uropathogen in a symptomatic patient. For catheterized specimens, ≥10^2 CFU/mL may be significant.
Yes. Possible reasons: recent antibiotic use, low colony count UTI, fastidious bacteria (like Ureaplasma), or non-infectious causes of symptoms.
To deliver a known volume of urine (e.g., 1 µL or 10 µL) onto the agar, allowing for the calculation of CFU/mL by counting colonies.
Blood Agar (BA): For overall growth and hemolysis. MacConkey Agar (MAC): Selective for Gram-negatives, differentiates lactose fermenters. CLED Agar: Supports both, prevents Proteus swarming.
Lactose fermenters (LF): Pink colonies (e.g., E. coli, Klebsiella). Non-lactose fermenters (NLF): Colorless or clear colonies (e.g., Pseudomonas, Proteus).
Usually indicates contamination during collection. The lab typically does not proceed with identification/susceptibility. A repeat sample is recommended.
Swarming is characteristic of Proteus mirabilis (and some other Proteus/Providencia species). It can complicate colony counting.
Indole positive, Methyl Red positive, Voges-Proskauer negative, Citrate negative (IMViC: ++–), Lactose fermenter, Motile.
By its resistance to the antibiotic Novobiocin (zone size < 16mm in disc diffusion). It's also a common uropathogen in young women.
A standardized AST method where filter paper discs impregnated with antibiotics are placed on a Mueller-Hinton agar plate lawned with the test organism. The zone of inhibition is measured and interpreted as Susceptible, Intermediate, or Resistant using CLSI/EUCAST tables.
Minimum Inhibitory Concentration. The lowest concentration of an antibiotic that visibly inhibits the growth of a bacterium. It’s a quantitative measure of susceptibility.
Extended-Spectrum Beta-Lactamase. An enzyme produced by some Gram-negatives (e.g., E. coli, Klebsiella) that confers resistance to penicillins, cephalosporins, and aztreonam. ESBL producers are often multidrug-resistant.
Methicillin-Resistant Staphylococcus aureus. Resistant to all beta-lactams (penicillins, cephalosporins, carbapenems). Lab detection often uses a cefoxitin disc screen.
Vancomycin-Resistant Enterococcus (usually E. faecium or E. faecalis). An important nosocomial pathogen with limited treatment options.
To screen for asymptomatic bacteriuria in high-risk groups: pregnant women (to prevent pyelonephritis), patients undergoing urologic procedures, and renal transplant recipients.
Funguria (Candiduria). Common in patients with indwelling catheters, diabetes, or on broad-spectrum antibiotics. May represent colonization or true infection.
Matrix-Assisted Laser Desorption/Ionization-Time of Flight Mass Spectrometry allows for rapid and accurate identification of bacteria and yeast directly from colonies, often in minutes, replacing traditional biochemical methods.
Cannot detect all pathogens (e.g., intracellular bacteria like Chlamydia), time-consuming (24-48h), contamination risk, and may be negative if antibiotics were taken recently.
Urease breaks down urea into ammonia, alkalinizing the urine. This can lead to the precipitation of magnesium ammonium phosphate (struvite) and calcium phosphate, forming kidney stones.
Even low colony counts (≥10^2 CFU/mL) can be significant in catheter-associated UTIs (CAUTI).
PYR (pyrrolidonyl arylamidase) test is positive for Enterococcus and Streptococcus pyogenes (Group A Strep), helping to identify them.