Gram-Positive Bacteria: Structure, Pathogenesis & Clinical Diagnostics
A complete clinical microbiology guide to the most significant Gram-Positive bacteria examples, including cell wall architecture, culture media, biochemical identification, and the rise of drug-resistant Gram-Positive infections.

Complete Taxonomy & Complex Nomenclature
Understanding Gram-Positive Evolutionary Lineages
In evolutionary microbiology, Gram-positive bacteria are often classified as Monoderms (single membrane with a thick cell wall). Taxonomically, they are divided into two massive clades based on the Guanine-Cytosine (G+C) content of their DNA:
- Low G+C Content (Phylum Bacillota / Firmicutes): Includes the most common medical pathogens such as Staphylococcus, Streptococcus, Enterococcus, Bacillus, and Clostridium. They typically have less than 50% G+C in their genomes.
- High G+C Content (Phylum Actinomycetota / Actinobacteria): Includes highly complex, often branching bacteria such as Mycobacterium (acid-fast), Corynebacterium, Nocardia, and Actinomyces. They have greater than 50% G+C content.
The Monoderm Cell Wall Architecture
Unlike their Gram-negative counterparts, Gram-positive bacteria possess a highly distinct, rigid, and thick cell envelope. They lack an outer lipid membrane entirely.
1. Thick Peptidoglycan Layer
The hallmark of Gram-positive bacteria is a massive, multi-layered shield of peptidoglycan (murein) that is 20 to 80 nanometers thick (comprising 50-90% of the cell envelope). It consists of alternating NAG and NAM sugars cross-linked by peptide bridges, providing immense physical strength and preventing osmotic lysis.
2. Teichoic & Lipoteichoic Acids
Unique to Gram-positive bacteria, these are water-soluble anionic polymers embedded directly into the peptidoglycan. Lipoteichoic acids (LTA) are anchored to the underlying plasma membrane. They regulate cell division, dictate cell shape, mediate adherence to host tissues, and serve as major antigenic determinants that trigger host immunity.
3. Endospore Formation
A survival mechanism exclusive to certain Gram-positive genera (primarily Bacillus and Clostridium). When faced with nutrient deprivation, these bacteria can condense their DNA into highly resilient, dormant spores that survive boiling, desiccation, UV radiation, and chemical disinfectants for decades.
The Gram Stain Principle: Why They Stain Purple
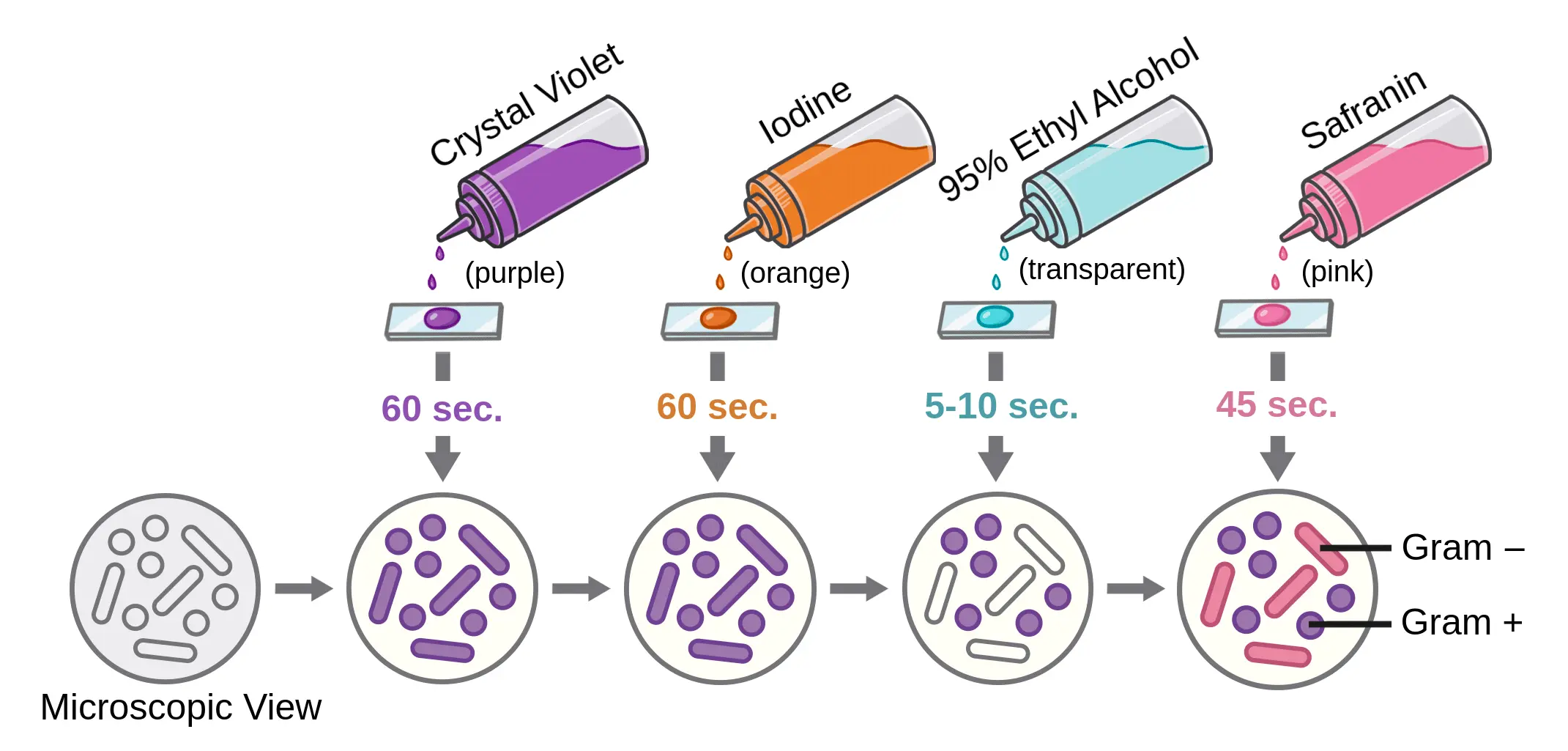
| Staining Step | Reagent Used | Effect on Gram-Positive Bacteria | Reasoning |
|---|---|---|---|
| 1. Primary Stain | Crystal Violet | Purple | The basic dye easily penetrates the thick cell wall. |
| 2. Mordant | Gram’s Iodine | Deep Purple | Iodine binds to crystal violet, forming large CV-I complexes. |
| 3. Decolorizer | Alcohol / Acetone | Remains Purple | Alcohol dehydrates the thick peptidoglycan, closing pores and trapping the dye inside. |
| 4. Counterstain | Safranin / Carbol Fuchsin | Remains Purple | The cell is already saturated with the darker purple dye; the lighter pink dye is masked. |
Colony Morphology & Essential Culture Media
Clinical isolation relies heavily on enriched and selective media to isolate Gram-positive organisms from mixed clinical specimens. The table below outlines 8 critical culture media used extensively for Gram-positives.
| Culture Media | Primary Use / Target Organisms | Key Morphology & Interpretation |
|---|---|---|
| Blood Agar Plate (BAP) | General enriched media; differentiates Streptococcus & Staphylococcus. | Demonstrates Hemolysis: Beta (clear zone, total lysis), Alpha (green zone, partial lysis), or Gamma (no lysis). |
| Mannitol Salt Agar (MSA) | Highly selective (7.5% salt) for Staphylococcus. Differential for S. aureus. | S. aureus ferments mannitol, turning the red agar to Bright Yellow. Coagulase-negative Staph (CoNS) remain red/pink. |
| Columbia CNA Agar | Selective isolation of Gram-positive cocci from mixed flora. | Contains Colistin & Nalidixic Acid to suppress Gram-negatives. Gram-positives grow well and exhibit true hemolysis. |
| Phenylethyl Alcohol Agar (PEA) | Selective for Gram-positive bacteria (especially anaerobes). | Phenylethyl alcohol dissolves the outer membrane of Gram-negatives. Streptococcus and Staphylococcus grow normally. |
| Bile Esculin Agar (BEA) | Selective & differential for Enterococcus and Group D Streptococcus. | Organisms that hydrolyze esculin in the presence of bile produce esculetin, which reacts with iron to turn the agar Dark Black. |
| Egg Yolk Agar (EYA) | Isolation and differentiation of Clostridium and Bacillus species. | Detects Lecithinase (opaque white halo around colony) and Lipase (iridescent “oil on water” sheen). |
| Tinsdale Agar / Tellurite Agar | Highly selective for Corynebacterium diphtheriae. | Potassium tellurite is reduced by the bacteria, producing distinctive Black colonies with a brown halo. |
| Chocolate Agar (CHOC) | Fastidious organisms, including heavily encapsulated Streptococcus pneumoniae. | Lysed red blood cells provide Factor V and X. S. pneumoniae typically forms glistening, mucoid, umbilicated (dimpled) colonies. |
Biochemical Identification Summary
Distinguishing between the major Gram-positive cocci and bacilli is accomplished through rapid benchtop biochemical tests.
| Biochemical Test | Positive Organisms (Examples) | Key Mechanism / Utility |
|---|---|---|
| Catalase Test | Staphylococcus, Micrococcus, Bacillus | Enzyme converts H₂O₂ into water and O₂ gas (bubbles). Crucial first step: Separates Staphylococcus (+) from Streptococcus (-). |
| Coagulase Test | Staphylococcus aureus | Enzyme converts fibrinogen to fibrin, causing plasma to clot. Separates S. aureus (+) from all other CoNS (-). |
| Optochin Susceptibility (P Disc) | Streptococcus pneumoniae | Zone of inhibition around the disc. Differentiates S. pneumoniae (Susceptible) from Viridans group Strep (Resistant). |
| Bacitracin Susceptibility (A Disc) | Streptococcus pyogenes (Group A) | Zone of inhibition. Separates Group A Strep (Susceptible) from Group B S. agalactiae (Resistant). |
| CAMP Test | Streptococcus agalactiae (Group B) | Produces CAMP factor that acts synergistically with S. aureus beta-toxin to create an “arrowhead” zone of enhanced hemolysis. |
| PYR Test | Enterococcus spp., S. pyogenes | Detects pyrrolidonyl arylamidase enzyme (turns bright red/pink). Rapid test to confirm Group A Strep or Enterococcus. |
Major Gram-Positive Bacteria List & Examples
1. The Staphylococci (Clusters)
Key Members: Staphylococcus aureus, S. epidermidis, S. saprophyticus.
Profile: Catalase-positive, non-motile cocci arranging in grape-like clusters. S. aureus is a highly virulent coagulase-positive pathogen notorious for abscesses and toxin-mediated diseases. S. epidermidis is a major biofilm producer on indwelling catheters. S. saprophyticus causes UTIs in young females.
2. The Streptococci (Chains)
Key Members: Streptococcus pyogenes, S. pneumoniae, S. agalactiae.
Profile: Catalase-negative cocci arranging in chains or pairs. Classified by Lancefield groupings (A, B, C, etc.) and hemolysis patterns. S. pyogenes (Group A) causes Strep throat, necrotizing fasciitis, and Rheumatic fever. S. pneumoniae (lancet-shaped diplococci) is the leading cause of community-acquired pneumonia and adult meningitis.
3. Spore-Forming Bacilli
Key Members: Bacillus anthracis, Clostridium tetani, C. botulinum, C. difficile.
Profile: Large Gram-positive rods capable of forming endospores. Bacillus are aerobic/facultative, causing Anthrax (boxcar-shaped rods) and food poisoning (B. cereus). Clostridium are strict anaerobes notorious for producing the most potent neurotoxins and enterotoxins known to medicine.
4. Non-Spore-Forming Bacilli
Key Members: Listeria monocytogenes, Corynebacterium diphtheriae.
Profile: Listeria is a psychrotroph (grows at refrigerator temperatures) causing severe meningitis in neonates and pregnant women, exhibiting unique “tumbling motility.” Corynebacterium form “club-shaped” arrangements (V/L shapes or palisades) and cause severe respiratory diphtheria via pseudomembrane formation.
5. Enterococcus Species
Key Members: Enterococcus faecalis, E. faecium.
Profile: Once classified as Group D Streptococcus, these hardy gastrointestinal commensals are highly tolerant to bile and salt. They are significant causes of nosocomial UTIs, biliary tract infections, and endocarditis, and are notorious for high levels of intrinsic and acquired antibiotic resistance (VRE).
6. The Actinomycetes
Key Members: Actinomyces israelii, Nocardia asteroides.
Profile: Gram-positive bacteria that form branching, filamentous networks resembling fungi. Actinomyces causes chronic cervicofacial abscesses with “sulfur granules.” Nocardia is weakly acid-fast and primarily causes pulmonary and cutaneous infections in immunocompromised patients.
Differences Between Gram Negative and Gram Positive Bacteria
| Characteristic / Feature | Gram-Positive Bacteria | Gram-Negative Bacteria |
|---|---|---|
| 1. Gram Stain Color | Purple / Violet | Pink / Red |
| 2. Peptidoglycan Layer | Thick (20-80 nm); heavily cross-linked. | Thin (1-3 nm); loosely cross-linked. |
| 3. Outer Membrane | Absent (Monoderms). | Present (Diderms; lipid bilayer). |
| 4. Lipopolysaccharide (LPS) | Absent. | Present in the outer leaflet (Endotoxin). |
| 5. Teichoic Acids | Present (Wall teichoic & lipoteichoic acids). | Absent entirely. |
| 6. Periplasmic Space | Absent or very narrow/indistinct. | Present (large, well-defined space containing enzymes). |
| 7. Lipid Content of Cell Wall | Low (1-4%). | High (11-22%) due to the outer membrane. |
| 8. Porin Channels | Absent. | Present in the outer membrane for selective transport. |
| 9. Flagellar Basal Body | Two rings (S and M rings). | Four rings (L, P, S, and M rings). |
| 10. Toxin Production | Primarily Exotoxins (actively secreted). | Primarily Endotoxins (Lipid A), though some exotoxins exist. |
| 11. Susceptibility to Penicillin | Generally High (unless carrying resistance genes). | Generally Low (outer membrane blocks entry). |
| 12. Susceptibility to Lysozyme | Highly susceptible (cleaves thick peptidoglycan). | Resistant (protected by outer membrane). |
| 13. Resistance to Physical Disruption | High (very strong, rigid cell wall). | Low (fragile cell wall). |
| 14. Resistance to Drying/Desiccation | High (survives well on dry surfaces). | Low (prone to rapid drying). |
| 15. Spore Formation capability | Present in specific genera (e.g., Bacillus, Clostridium). | Absent entirely. |
Similarities Between Gram Negative and Gram Positive Bacteria
| Shared Feature | Biological Explanation & Details |
|---|---|
| 1. Cellular Classification | Both are strictly prokaryotic cells, lacking a true nucleus and membrane-bound organelles. |
| 2. Genetic Architecture | Both possess a primary nucleoid region containing a circular, double-stranded DNA chromosome. |
| 3. Plasmid Capability | Both frequently harbor extrachromosomal DNA plasmids that carry antibiotic resistance and virulence genes. |
| 4. Protein Synthesis Apparatus | Both utilize 70S ribosomes (comprising 50S and 30S subunits) for mRNA translation into proteins. |
| 5. Cytoplasmic Membrane | Both feature a selectively permeable inner phospholipid bilayer that encloses the cytoplasm. |
| 6. Cytoplasm Presence | Both contain an aqueous, gel-like cytoplasm where primary metabolic and enzymatic reactions occur. |
| 7. Peptidoglycan Presence | Both utilize peptidoglycan (murein) composed of NAG and NAM to maintain structural integrity and shape. |
| 8. Method of Reproduction | Both divide and multiply primarily through asexual binary fission. |
| 9. Surface Motility Appendages | Both classifications contain species capable of producing flagella for chemotaxis and motility. |
| 10. Adherence Structures | Both can produce pili and fimbriae to assist in attaching to host tissues or environmental surfaces. |
| 11. Capsular Polysaccharides | Both can secrete thick extracellular polysaccharide capsules to evade host phagocytosis and complement. |
| 12. Biofilm Production | Both readily secrete extracellular polymeric substances (EPS) to form highly resistant biofilms on medical devices. |
| 13. Core Metabolic Pathways | Both rely on universal core energy pathways such as glycolysis and the TCA cycle (when aerobic). |
| 14. Initial Gram Stain Reaction | Both cell types actively take up the primary Crystal Violet stain and form internal Iodine complexes initially. |
| 15. Susceptibility to Antimicrobials | Both possess shared molecular targets for antibiotics (e.g., DNA gyrase, RNA polymerase, 30S/50S ribosomes). |
Major Gram-Positive Infections & Pathogenesis
Because of their diverse array of exotoxins and adherence factors, Gram-Positive infections account for a massive percentage of worldwide human morbidity, ranging from mild skin conditions to rapidly fatal systemic toxemias.
A. Skin and Soft Tissue Infections (SSTIs)
Staphylococcus aureus and Streptococcus pyogenes are the undisputed kings of the skin. They cause impetigo, folliculitis, cellulitis, severe carbuncles, and potentially limb-threatening Necrotizing Fasciitis (“flesh-eating disease”). Their surface proteins (like Protein A in Staph and M Protein in Strep) strongly inhibit phagocytosis by white blood cells.
B. Respiratory & Pulmonary Infections
The upper respiratory tract is commonly afflicted by Streptococcus pyogenes (Bacterial Pharyngitis / Strep Throat) and Corynebacterium diphtheriae. In the lower respiratory tract, Streptococcus pneumoniae is the leading cause of lobar pneumonia, utilizing a thick capsule to avoid immune clearance within the alveoli.
C. Toxin-Mediated Systemic Syndromes
- Toxic Shock Syndrome (TSS): Caused by Superantigens released by S. aureus or S. pyogenes, creating a massive cytokine storm.
- Tetanus & Botulism: Caused by Clostridium neurotoxins. Tetanospasmin causes spastic paralysis (lockjaw), while Botulinum toxin causes flaccid paralysis.
- Pseudomembranous Colitis: Caused by Exotoxins A and B from Clostridium difficile, usually following extensive broad-spectrum antibiotic usage.
Treatment Strategies & Antimicrobial Resistance (AMR)
Standard Therapeutics
- Penicillins & Cephalosporins: Target Penicillin-Binding Proteins (PBPs) to halt cell wall synthesis. Highly effective against S. pyogenes.
- Macrolides: Azithromycin and Erythromycin inhibit the 50S ribosomal subunit. Often used for respiratory Strep infections in penicillin-allergic patients.
- Clindamycin: Excellent tissue penetration; effectively halts exotoxin production in severe Staph/Strep SSTIs.
Resistant “Superbugs” & Last-Resort Drugs
When resistance occurs, therapeutics must shift to highly specific or potent alternatives:
- MRSA (Methicillin-Resistant S. aureus): Harbors the mecA gene, altering the PBP target (PBP2a). Treated primarily with IV Vancomycin or Daptomycin.
- VRE (Vancomycin-Resistant Enterococcus): Alters the cell wall peptide precursor (D-ala-D-ala to D-ala-D-lac) so Vancomycin cannot bind. Treated with Linezolid or Daptomycin.
- PRSP (Penicillin-Resistant S. pneumoniae): Acquired through altered PBPs via transformation. Often requires highly dosed 3rd generation cephalosporins or respiratory fluoroquinolones.
Rapid Review: MLS / ASCP Board Pearls
The Catalase Branch
If you see a Gram-positive coccus, immediately check Catalase. Positive (Bubbles) = Staphylococci/Micrococci. Negative = Streptococci/Enterococci.
Coagulase Dictates Staph
Among the Catalase-positive Staph, Coagulase Positive almost always equals Staphylococcus aureus. All others are lumped into CoNS (Coagulase-Negative Staph).
Spore-Forming Rods
If you have a Gram-Positive Rod (Bacillus) that forms spores, it is either Bacillus (aerobic/catalase-positive) or Clostridium (strict anaerobic/catalase-negative).
Hemolysis & Discs
Beta-hemolytic Strep? Check Bacitracin (Group A is sensitive). Alpha-hemolytic? Check Optochin (S. pneumoniae is sensitive, Viridans is resistant).
FAQs:
What are Gram-positive bacteria?
Gram-positive bacteria are a group of bacteria that have a thick peptidoglycan layer in their cell wall, which stains purple when exposed to the Gram stain.
What are some examples of Gram-positive bacteria?
Examples of Gram-positive bacteria include Staphylococcus aureus, Streptococcus pneumoniae, and Bacillus subtilis.
What are the characteristics of Gram-positive bacteria?
Characteristics of Gram-positive bacteria include a thick peptidoglycan layer in their cell wall, lack of an outer membrane, and the ability to form endospores.
What are the ecological roles of Gram-positive bacteria?
Gram-positive bacteria play a variety of ecological roles, including nitrogen fixation, decomposition, and fermentation.
What are the industrial applications of Gram-positive bacteria?
Gram-positive bacteria are used in a variety of industrial applications, including food production, bioremediation, and the production of antibiotics and enzymes.
How are Gram-positive bacterial infections diagnosed and treated?
Gram-positive bacterial infections can be diagnosed through various laboratory tests, and are typically treated with antibiotics.
What are some common diseases caused by Gram-positive bacteria?
Common diseases caused by Gram-positive bacteria include pneumonia, sepsis, and skin infections.
What are some virulence factors of Gram-positive bacteria?
Virulence factors of Gram-positive bacteria include toxins, adhesins, and capsule polysaccharides.
What is antibiotic resistance in Gram-positive bacteria?
Antibiotic resistance in Gram-positive bacteria occurs when the bacteria develop the ability to resist the effects of antibiotics, making them difficult to treat.
What are some mechanisms of antibiotic resistance in Gram-positive bacteria?
Mechanisms of antibiotic resistance in Gram-positive bacteria include the production of antibiotic-degrading enzymes, efflux pumps, and modification of antibiotic targets.
How can Gram-positive bacterial infections be prevented and controlled?
Prevention and control of Gram-positive bacterial infections can be achieved through measures such as good hygiene, vaccination, and the appropriate use of antibiotics.
What are some future directions in Gram-positive bacterial research?
Future research in Gram-positive bacterial studies will likely focus on understanding bacterial virulence, mechanisms of antibiotic resistance, and the microbiome, with the goal of developing new treatments and preventative measures to combat bacterial infections.
What is the role of Gram-positive bacteria in the human microbiome?
Gram-positive bacteria are a major component of the human microbiome, and are thought to play important roles in digestion, immune function, and protection against pathogens.
What are some methods for studying Gram-positive bacteria?
Methods for studying Gram-positive bacteria include culturing the bacteria in the lab, genetic manipulation, and microscopy.
What are some applications of CRISPR-Cas in Gram-positive bacterial research?
CRISPR-Cas can be used to study and manipulate Gram-positive bacterial genomes, which can help researchers better understand the virulence factors and regulatory networks that are important for bacterial survival and pathogenesis.
What are biofilms, and how do they relate to Gram-positive bacteria?
Biofilms are communities of bacteria that form on surfaces and are held together by a matrix of extracellular polymeric substances. Gram-positive bacteria are known to form biofilms, which can be difficult to treat with antibiotics.
How do Gram-positive bacteria differ from Gram-negative bacteria?
Gram-positive bacteria have a thick peptidoglycan layer in their cell wall, lack an outer membrane, and stain purple when exposed to the Gram stain. Gram-negative bacteria have a thinner peptidoglycan layer, an outer membrane, and stain pink when exposed to the Gram stain.