Gram-Negative Bacteria: Structure, Pathogenesis & Antimicrobial Resistance
The complete encyclopedic reference for Clinical Microbiology covering cell wall biology, Gram staining principles, lab diagnostics, and the growing global crisis of Multi-Drug Resistant (MDR) infections.

Classification & Important Medical Genera
| Enterobacteriaceae (Bacilli) | Escherichia coli, Klebsiella, Salmonella, Shigella, Proteus |
|---|---|
| Non-Fermenting Bacilli | Pseudomonas aeruginosa, Acinetobacter baumannii |
| Gram-Negative Cocci | Neisseria gonorrhoeae, Neisseria meningitidis, Moraxella |
| Fastidious Coccobacilli | Haemophilus influenzae, Bordetella pertussis, Brucella |
| Curved / Spiral Rods | Vibrio cholerae, Campylobacter jejuni, Helicobacter pylori |
| Obligate Intracellular | Chlamydia trachomatis, Rickettsia rickettsii |
Understanding Gram-Negative Taxonomy
Gram-negative bacteria represent a massive portion of the bacterial domain. In evolutionary microbiology, they are often referred to as “Diderms” because they possess two cellular membranes (an inner cytoplasmic membrane and an outer lipid membrane), distinguishing them from Gram-positive “Monoderms.” This comprehensive Gram-negative bacteria list showcases their immense structural and functional diversity.
- Shape Variations: While the majority of medically significant Gram-negatives are bacilli (rods), they also appear as cocci (spheres), coccobacilli (short rods), and curved/helical forms.
- Environmental Adaptability: Found in every ecosystem on Earth, from human mucosal surfaces to deep-sea thermal vents, due to their highly protective outer membrane.
The Diderm Cell Wall Structure
The defining characteristic of Gram-negative bacteria is their highly complex cell envelope. Unlike Gram-positive bacteria which have a single thick layer of peptidoglycan, Gram-negatives feature a thin layer sandwiched between two distinct lipid bilayers.
1. The Outer Membrane & LPS
A unique lipid bilayer serving as a formidable permeability barrier against large molecules, lysozymes, and many antibiotics. The outer leaflet contains Lipopolysaccharide (LPS). Porins are protein channels in the outer membrane that allow passive diffusion of small hydrophilic molecules, including some antibiotics. Notably, Lipid A is the toxic component of LPS responsible for fever, shock, and severe systemic inflammation.
2. Periplasmic Space
The gel-like matrix located between the inner cytoplasmic membrane and the outer membrane. This compartment is densely packed with hydrolytic enzymes, nutrient-binding proteins, and crucially, Beta-Lactamases—enzymes that destroy antibiotics before they can reach the cell wall machinery.
3. Thin Peptidoglycan Layer
A very thin layer (approx. 1-3 nm) of murein (peptidoglycan) that provides structural integrity and shape. Because it is so thin and sparsely cross-linked, it fails to retain the primary crystal violet stain during the Gram staining procedure.
The Gram Stain Principle: Gram-Negative vs Gram-Positive
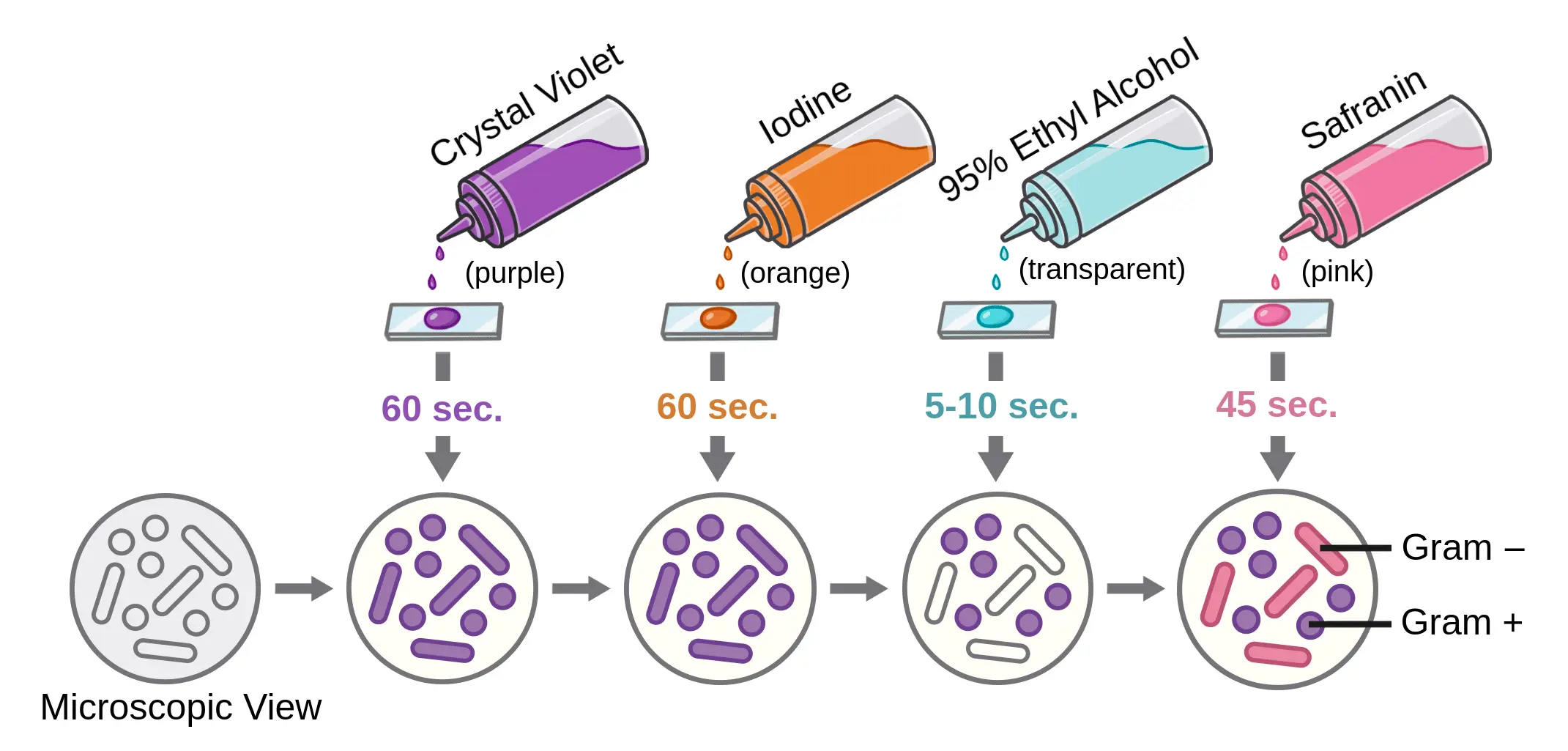
| Staining Step | Reagent Used | Effect on Gram-Negative Bacteria | Effect on Gram-Positive Bacteria |
|---|---|---|---|
| 1. Primary Stain | Crystal Violet | Stain purple | Stain purple |
| 2. Mordant | Gram’s Iodine | Complexes form; remain purple | Complexes form strongly; remain purple |
| 3. Decolorizer | Alcohol / Acetone | Colorless (Outer membrane dissolves, dye escapes) | Remain Purple (Thick PG dehydrates, trapping dye) |
| 4. Counterstain | Safranin or Carbol Fuchsin | Stain Pink/Red | Remain Purple (Pink dye is masked by darker purple) |
Differences Between Gram-Negative and Gram-Positive Bacteria
| Feature | Gram-Negative Bacteria | Gram-Positive Bacteria |
|---|---|---|
| 1. Gram Stain Color | Pink / Red | Purple / Blue |
| 2. Peptidoglycan Layer | Thin (approx. 1-3 nm) | Thick (approx. 20-80 nm) |
| 3. Outer Membrane | Present (Double membrane / Diderms) | Absent (Single membrane / Monoderms) |
| 4. Lipopolysaccharide (LPS) | Present (Source of Endotoxin/Lipid A) | Absent |
| 5. Teichoic Acids | Absent | Present (Lipoteichoic and wall teichoic acids) |
| 6. Periplasmic Space | Present (Well-defined and large) | Absent or very narrow |
| 7. Lipid Content | High (11-22% due to outer membrane) | Low (1-4%) |
| 8. Porin Proteins | Present in the outer membrane | Absent |
| 9. Flagellar Basal Body | Four rings (L, P, S, M) | Two rings (S, M) |
| 10. Toxin Production | Primarily Endotoxins (some exotoxins) | Primarily Exotoxins |
| 11. Susceptibility to Penicillin | Generally low (outer membrane acts as a barrier) | Generally high |
| 12. Susceptibility to Lysozyme | Low resistance | High resistance (unless treated beforehand) |
| 13. Resistance to Physical Disruption | Low (fragile cell wall) | High (sturdy, thick peptidoglycan) |
| 14. Resistance to Drying | Low (prone to desiccation) | High (survives well in dry environments) |
| 15. Mesosomes | Less prominent | More prominent |
Similarities Between Gram-Negative and Gram-Positive Bacteria
| Feature / Characteristic | Shared Biological Aspect |
|---|---|
| 1. Cell Type | Both are prokaryotic cells, lacking a true nucleus and membrane-bound organelles. |
| 2. Genetic Material | Both possess a primary genome composed of double-stranded, circular DNA localized in a nucleoid region. |
| 3. Extrachromosomal DNA | Both can harbor plasmids, enabling horizontal gene transfer and antimicrobial resistance. |
| 4. Ribosomes | Both contain 70S ribosomes (consisting of 50S and 30S subunits) responsible for protein synthesis. |
| 5. Cytoplasmic Membrane | Both have a selectively permeable inner phospholipid bilayer (plasma membrane). |
| 6. Cytoplasm | Both contain a gel-like cytoplasm where crucial cellular enzymatic reactions take place. |
| 7. Peptidoglycan Presence | Both use peptidoglycan (murein) networks to maintain their structural shape, albeit in different thicknesses. |
| 8. Reproduction | Both reproduce primarily through asexual reproduction via binary fission. |
| 9. Surface Appendages | Both can possess flagella for motility and pili/fimbriae for cellular adhesion. |
| 10. Pathogenicity | Both classifications include significant human opportunistic and obligate pathogens. |
| 11. Immune Evasion | Both can produce extracellular polysaccharide capsules to evade host phagocytosis. |
| 12. Biofilm Formation | Both are fully capable of secreting extracellular polymeric substances to form protective biofilms. |
| 13. Core Metabolism | Both utilize universal, core metabolic pathways such as glycolysis for energy production. |
| 14. Gram Stain Uptake | Both initially take up the primary crystal violet stain and form iodine complexes before the decolorization step. |
| 15. Antibiotic Targets | Both share common foundational targets for antibiotics (e.g., ribosomes for macrolides, DNA replication enzymes for fluoroquinolones). |
Major Gram-Negative Bacteria Examples & Pathogens
1. Enterobacteriaceae (The Enterics)
Key Members: Escherichia coli, Klebsiella pneumoniae, Proteus mirabilis.
Clinical: The most common causes of Urinary Tract Infections (UTIs), intra-abdominal infections, and healthcare-associated pneumonias. Most Enterobacteriaceae are facultative anaerobes, allowing survival in both aerobic and anaerobic environments. Uropathogenic E. coli (UPEC) uses P-fimbriae to ascend the urinary tract and resist being flushed out by urine.
2. Pseudomonas aeruginosa
Profile: A ubiquitous, non-fermenting, strictly aerobic Gram-negative rod.
Clinical: An opportunistic powerhouse notorious for forming thick biofilms. It has intrinsic resistance to many drugs due to low outer membrane permeability and the presence of multiple efflux pumps. It is a leading cause of Ventilator-Associated Pneumonia (VAP) and produces a distinct green/blue pigment (pyocyanin).
3. Pathogenic Neisseria
Key Members: Neisseria meningitidis, Neisseria gonorrhoeae.
Clinical: Gram-negative diplococci (kidney-bean shaped). N. meningitidis causes rapidly fatal epidemic meningitis. Diagnosis includes Gram stain showing intracellular diplococci within neutrophils, growth on selective Thayer-Martin agar, and rapid oxidase positivity.
4. Gastrointestinal & Zoonotic
Key Members: Salmonella, Shigella, Campylobacter, Helicobacter.
Clinical: Primary agents of foodborne illness and dysentery. Helicobacter pylori uniquely survives extreme stomach acidity via urease production, causing peptic ulcers and gastric carcinoma. Vibrio cholerae causes massive watery “rice-water” diarrhea via the cholera toxin.
Mechanisms of Pathogenicity
A. Endotoxin (Lipid A) & Septic Shock
Unlike exotoxins which are actively secreted, Endotoxin is a structural component of the Gram-negative outer membrane. Endotoxin (Lipid A) is primarily released during cell lysis, but small amounts may also be shed during bacterial growth. Lipid A is the toxic component of LPS responsible for fever, shock, and inflammation. When released into the bloodstream, it hyper-activates macrophages via Toll-Like Receptor 4 (TLR4), causing a massive “cytokine storm” (IL-1, TNF-alpha). This leads to profound hypotension, Disseminated Intravascular Coagulation (DIC), and potentially fatal Septic Shock.
B. Pili (Fimbriae) & Adhesion
Hair-like protein appendages extending from the cell surface. They are essential for anchoring the bacteria to host tissues, preventing them from being washed away by urine or mucus. For example, P-fimbriae in Uropathogenic E. coli tightly bind to urothelial cells, causing pyelonephritis.
C. Type III Secretion Systems (T3SS)
A molecular “syringe” structurally similar to a flagellum, found in pathogens like Salmonella, Shigella, E. coli, and Pseudomonas. It directly injects bacterial effector proteins into the host cell cytoplasm, forcing the host cell to engulf the bacteria or undergo apoptosis.
D. Capsules & Biofilm Formation
Polysaccharide capsules (e.g., in K. pneumoniae, H. influenzae, N. meningitidis) hide antigenic targets from host antibodies and prevent phagocytosis. Many Gram-negatives also secrete an extracellular polymeric substance (EPS) to form Biofilms on catheters, prosthetic valves, and lungs, making them functionally impervious to immune cells and antibiotics.
The Antimicrobial Resistance (AMR) Crisis
| Resistance Mechanism | How it Works | Clinical Impact |
|---|---|---|
| Beta-Lactamases (ESBLs & Carbapenemases) | Enzymes secreted into the periplasmic space that hydrolyze the beta-lactam ring of antibiotics (Penicillins, Cephalosporins). | Renders entire classes of antibiotics useless. ESBL (Extended-Spectrum Beta-Lactamase) organisms require Carbapenems. CRE organisms destroy even Carbapenems. |
| Efflux Pumps | Transmembrane protein channels that actively pump antibiotics out of the bacterial cell faster than they can enter. | Confer multi-drug resistance (MDR) to diverse drug classes simultaneously, especially Tetracyclines and Fluoroquinolones. |
| Porin Channel Mutations | Alteration or loss of outer membrane porins (the “doors” used by water-soluble drugs to enter). | Antibiotics (like Imipenem in Pseudomonas) physically cannot reach their target sites inside the cell. |
Laboratory Diagnostics & Identification
Colony Morphology on Common Media
| Media | Colony Appearance | Interpretation |
|---|---|---|
| MacConkey (MAC) Agar | Pink (Lactose Fermenters) vs. Colorless (Non-Fermenters) | Differentiates rapid coliforms (E. coli, Klebsiella) from non-fermenters (Salmonella, Pseudomonas). |
| XLD Agar | Red colonies ± black center (H₂S) | Used primarily for recovering Salmonella and Shigella. |
| SS Agar | Colorless ± black center | Highly selective for Salmonella & Shigella from stool samples. |
| HE Agar | Green/blue ± black center | Hektoen Enteric agar isolates gastrointestinal pathogens. |
| Blood Agar | Variable (Large grey colonies, hemolysis may be present) | Non-selective; demonstrates beta-hemolysis often seen in E. coli strains. |
Biochemical Identification Summary
| Test | Positive Organisms (Examples) | Key Mechanism |
|---|---|---|
| Oxidase | Pseudomonas, Neisseria, Vibrio | Presence of cytochrome C oxidase. Separates them from Enterobacteriaceae (which are negative). |
| Indole | E. coli, Proteus vulgaris | Production of tryptophanase enzyme, converting tryptophan to indole (Red ring). |
| Urease | Proteus, Helicobacter, Klebsiella | Enzyme hydrolyzes urea to ammonia, turning the media bright pink (alkaline). |
| H₂S Production | Salmonella, Proteus, Citrobacter | Reduction of sulfur produces hydrogen sulfide gas (Black precipitate on media). |
Treatment Strategies for Gram-Negative Infections
Due to the protective outer membrane and extensive resistance mechanisms, treating severe Gram-negative infections requires specific antimicrobial classes that can successfully penetrate the periplasmic space or disrupt the cellular envelope.
Common Gram-Negative Antibiotics
- Beta-Lactam/Inhibitor Combos: Piperacillin/Tazobactam (Zosyn), Amoxicillin/Clavulanate.
- Cephalosporins: 3rd Generation (Ceftriaxone, Ceftazidime) and 4th Generation (Cefepime).
- Carbapenems: Meropenem, Imipenem (Often reserved for ESBL-producing bacteria).
- Aminoglycosides: Gentamicin, Tobramycin, Amikacin (Protein synthesis inhibitors).
- Fluoroquinolones: Ciprofloxacin, Levofloxacin (DNA gyrase inhibitors).
Last-Resort Therapeutics
For extensively drug-resistant (XDR) strains like CRE or pan-resistant Acinetobacter, treatment options are drastically limited.
- Polymyxins (Colistin): Acts like a detergent to disrupt the Gram-negative outer membrane. Highly nephrotoxic but often life-saving.
- Novel Combos: Ceftazidime/Avibactam (targets some carbapenemases).
- Tigecycline: A broad-spectrum bacteriostatic agent, though ineffective against Pseudomonas.
Rapid Review: MLS / ASCP Board Pearls
The Oxidase Rule
If you have a Gram-Negative Rod, immediately check Oxidase. Oxidase Negative → Enterobacteriaceae. Oxidase Positive → Think non-enterics (Pseudomonas, Vibrio, Campylobacter).
Lactose Fermenter Trick
Fast lactose fermenter = E. coli (produces flat, dry, bright pink colonies with a precipitated bile halo on MAC agar). Klebsiella is pink but highly mucoid.
H₂S Producers & Swarming
Salmonella & Proteus both produce black colonies on SS/XLD/HE. Differentiate them with Urease (Proteus is rapidly positive) and motility. Proteus mirabilis “swarms” over the entire agar plate.
Neisseria Hallmarks
Gram-Negative Diplococci (kidney beans) found inside Polymorphonuclear Leukocytes (PMNs) from urethral discharge is highly diagnostic for N. gonorrhoeae.
FAQs:
What are Gram-negative bacteria?
Gram-negative bacteria are a group of bacteria characterized by the staining properties of their cell walls, which do not retain the crystal violet dye used in the Gram staining method.
What are some examples of Gram-negative bacteria?
Some examples of Gram-negative bacteria include Escherichia coli, Salmonella, Pseudomonas aeruginosa, Neisseria gonorrhoeae, and Klebsiella pneumoniae.
What is the pathogenicity of Gram-negative bacteria?
Gram-negative bacteria can cause a wide range of diseases, including urinary tract infections, pneumonia, meningitis, and sepsis.
What are some virulence factors of Gram-negative bacteria?
Some virulence factors of Gram-negative bacteria include endotoxins, exotoxins, pili, flagella, and capsule.
How are Gram-negative bacterial infections diagnosed?
Gram-negative bacterial infections are typically diagnosed through a combination of clinical symptoms, laboratory tests such as blood cultures and urine cultures, and imaging studies.
How are Gram-negative bacterial infections treated?
Gram-negative bacterial infections are typically treated with antibiotics, although the choice of antibiotic may depend on the specific bacteria causing the infection and its antibiotic susceptibility.
What are some mechanisms of antibiotic resistance in Gram-negative bacteria?
Some mechanisms of antibiotic resistance in Gram-negative bacteria include efflux pumps, enzymatic degradation of antibiotics, and alteration of antibiotic targets.
What is the ecological importance of Gram-negative bacteria?
Gram-negative bacteria play important roles in various ecological processes, such as nutrient cycling, biodegradation of organic matter, and nitrogen fixation.
What is the industrial importance of Gram-negative bacteria?
Gram-negative bacteria are used in various industrial applications, such as production of antibiotics, bioremediation of pollutants, and bioprocessing of food and beverage products.
What is the role of Gram-negative bacteria in the human microbiome?
Gram-negative bacteria are an important component of the human microbiome, where they play important roles in host immune function, metabolism, and protection against pathogenic bacteria.
What are some challenges in Gram-negative bacterial research?
Some challenges in Gram-negative bacterial research include antibiotic resistance, pathogenesis, virulence, and development of new treatments.
What is the potential for phage therapy in Gram-negative bacterial infections?
Phage therapy, which involves using bacteriophages to target and kill specific bacteria, has shown promise as a potential treatment for Gram-negative bacterial infections, particularly in the context of antibiotic resistance.
What is the potential for immunotherapy in Gram-negative bacterial infections?
Immunotherapy, which involves using the immune system to target and eliminate specific bacteria, has shown promise as a potential treatment for Gram-negative bacterial infections, particularly in the context of antibiotic resistance.
What is the role of Gram-negative bacteria in foodborne illness?
Several Gram-negative bacteria, such as Salmonella and Escherichia coli, can cause foodborne illness through consumption of contaminated food products.
What is the role of Gram-negative bacteria in waterborne illness?
Several Gram-negative bacteria, such as Vibrio cholerae and Legionella pneumophila, can cause waterborne illness through consumption of contaminated water or inhalation of aerosolized water droplets.
How can we prevent and control Gram-negative bacterial infections?
Prevention and control of Gram-negative bacterial infections can involve measures such as good hygiene practices, appropriate use of antibiotics, vaccination, and public health measures such as surveillance and outbreak investigations.
References:
- Todar, K. (2012). Todar’s online textbook of bacteriology. Gram-negative bacteria. Retrieved from http://textbookofbacteriology.net/index.html
- Brooks, G. F., Carroll, K. C., Butel, J. S., Morse, S. A., & Mietzner, T. A. (2013). Jawetz, Melnick, & Adelberg’s Medical Microbiology. New York: McGraw-Hill Medical.
- Peleg, A. Y., Seifert, H., & Paterson, D. L. (2008). Acinetobacter baumannii: Emergence of a Successful Pathogen. Clinical microbiology reviews, 21(3), 538-582.
- Rafferty, S. (2001). Gram-negative bacteria. In Lederberg J. (eds) Encyclopedia of Microbiology (pp. 288-298). Academic Press.
- Silhavy, T. J., Kahne, D., & Walker, S. (2010). The bacterial cell envelope. Cold Spring Harbor perspectives in biology, 2(5), a000414.